Introduction
Plant cell walls
Pectins
Pectin methylesterases (PMEs)
Catalytic properties of PME
Pectin Methylesterase Inhibitor (PMEI)
PMEI structure
PME and PMEI interaction
Functions of PMEs and PMEIs in plant development
Seed germination
Organ formation in shoot apical meristem (SAM)
Wood development
Pollen formation and germination
Fruit development and ripening
Conclusion
Introduction
Unlike animal cells, plant cells possess outermost protected structures called cell walls. The cell wall plays a primary role in plant growth, development and response to environment stimuli. While the walls support the rigidity, strengths and protections for plant cells, it is still very dynamic and flexible to serve functions in cell divisions, elongations and differentiations required during plant development by a number of different cell wall hydrolases and esterases.
Pectin is the dominant cell wall polysaccharide in middle lamella and principal component in primary cell walls of dicots and none-grass monocots. Based on the substitution and side chain sugars, pectin is classified into four groups. Among them, homogalacturonan (HG) account for approximately 70-80% of total pectin in plant cell walls. Pectin is synthesized in the Golgi apparatus, secreted into the cell wall in a highly methyl esterified form. The highly methylesterified HG can be selectively demethylesterified by PME (Pelloux et al., 2007). PME catalyses the hydrolysis of methyl-ester bonds at C6 of methyl-D-galactopyranosyl residues of pectin, forming negatively charged polymer groups and methanol. After demethylesterification, de-methylesterified pectin can form Ca2+-pectate cross-link complexes, stiffening cell walls or can be a target for pectin-degrading enzymes such as polygalacturonases that subsequently affect the texture and rigidity of plant cell walls. Thus, the cell wall structures and properties are strongly affected by the degree and distribution of pectin methyl esterification. Recent studies also demonstrated that fine-tuning of pectin methylesterification status plays an important role in diverse plant developmental processes and stress responses (Pelloux et al., 2007; Wolf et al., 2009).
Plant PMEs have multiple isoforms encoded by large multigene families in both monocots and dicots. For instance, there are 66 PMEs in Arabidopsis thaliana, 79 in Solanum lycopersicum, 89 in Poplus trichocarpa, 105 in Linum usitatissimum and 43 in Oryza sativa (Jeong et al., 2015; Pelloux et al., 2007; Pinzón-Latorre and Deyholos, 2013; Vandevenne et al., 2009). The expression of PME isoforms were precisely regulated at transcriptional level during developmental stages and in specific tissues. In addition, the PME activity also be regulated by specific proteinaeous inhibitors, PMEI, firstly discovered in kiwi (Actinidia deliciosa) (Balestrieri et al., 1990; Giovane et al., 1995; Giovane et al., 2004). PMEIs typically inhibit plant specific PMEs instead of microbial PMEs (Giovane et al., 2004; Jolie et al., 2010). Although grass species contain a low level of pectin (2-10%), they also possess large multigene families of PME and PMEI, such as Oryza sativa (43 PMEs and 49 PMEIs) and Brachypodium distachyon (29 PMEs and 38 PMEIs) suggesting the methylesterification of pectin is probably involved in plant developments and defenses as observation in dicot plants (Nguyen et al., 2016).
Plant cell walls
The plant cell wall located outside of the plasma membrane is composed of polysaccharides, protein and phenolic compounds (Cosgrove, 2005; Darvill et al., 1980; Fry, 1988; Scheller and Ulvskov, 2010). The composition and distribution of cell wall polysaccharides are different depending on the species and tissues. Basically, the cell wall polysaccharides have been classified into three group: cellulose, hemicellulose and pectin. Major functional roles of plant cell walls include mechanical support, protoplast protection, and determination of final cell shapes (Levesque-Tremblay et al., 2015a).
The cell walls are divided into two types, primary and secondary cell walls: the primary cell walls are the walls surrounding growing cells and present in all plant cells, secondary cell walls are found in some specialized cells including xylem and fiber cell types after cessation of primary cell wall growth, making the walls stronger. In addition, there are also structures in cell walls named middle lamella that cements the walls of neighbor’s cells together to render plant cells stability. The middle lamella is called as pectin layer because the main cell wall polysaccharide is pectins in forms of calcium and magnesium pectates. When a plant cell divides, a middle lamella and a primary cell wall is firstly established.
The primary cell walls are pliant, thin and highly hydrated structures composed of 15-40% cellulose, 30-50% pectins, 20-30% hemicelluloses, and proteins (Cosgrove, 2005; Cosgrove and Jarvis, 2007; Darvill et al., 1980). Cellulose is composed of unbranched (1, 4)-linked β-D-glucan chains which interact with each other to form cellulose microfibrils via hydrogen bonds. Due to the rigidity and strength, cellulose microfibrils is referred to as the cell wall skeleton (Fry, 2004). Hemicelluloses characterized by having a backbone of β-1,4-linked sugars are a heterogeneous group of polysaccharides including xyloglucans, xylans, mannans and glucomannans, and β-(1, 3, 1, 4) mixed-glucans (Scheller and Ulvskov, 2010). They are chemically bonded to the cellulose microfibrils. Beside these mainly polysaccharides, there are some structural and enzymatic proteins were detected in primary cell wall (Cosgrove, 2005). The secondary cell walls are thick layers developed between the primary cell walls and plasma membrane in plant tissues ceased growing. This wall is generally not as dynamic and malleable as the primary cell wall. They strengthen and provide the rigidity for plant cell walls. The secondary cell walls contain cellulose, hemicellulose, pectins and lignin, which strengthens and waterproofs the cell wall. However, this composition is variable depending on the differentiated cell types (Keegstra, 2010).
Pectins
Pectins are abundant in walls in growing and dividing cells (primary cell walls) and in the middle lamella. They present approximately 35% and 2-10% in cell wall compositions of dicots as well as non-grass monocots and grass plants, respectively (Ochoa-Villarreal et al., 2012; Ridley et al., 2001). Pectins are thought to perform a variety of biological functions including cell wall porosity, cell expansion, regulation of cell adhesion, plant morphogenesis, pollen tube growth, leaf abscission, fruit development, seed hydration, providing charged surfaces for modulating the wall pH, and serve as signalling molecules in pathogen recognition and plant growth (Mohnen, 2008).
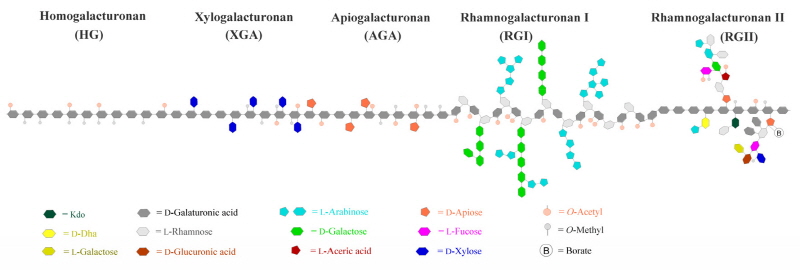
Pectins are a family of covalently linked galacturonic acid (GalUA)-rich polysaccharides including homogalac-turonan (HG), rhamnogalacturonan I (RGI), rhamnogalcturonan II (RGII) xylogalcturonan (XGA) and apiogalacturonan (AGA) (Harholt et al., 2010) (Fig. 1). Among pectic polysaccharides, HG is the major component, which constitutes approximately 60% of pectins, RGI constitutes 20-35%, while XGA and RGII are minor components accounting for less than 10% of pectins (Mohnen, 2008; Zandleven et al., 2007). AGA is a specific pectin found in aquatic plants such as duckweeds and marine seagrases (Ochoa-Villarreal et al., 2012). HG is unbranched homopolymer chains of α-1,4-linked D-GalUA. The backbone of GalUA can be attached additional xylose and apiose side chains to form XGA and AGA, respectively. RGII can be formed by adding clusters of complex side chains composed of 12 types of glycosyl residues linked together by at least 22 different glycosidic bonds on the GalUA chains. RGI is not built on pure GalUA backbone instead of a polymer with a backbone structure of (α-1,4-D-GalA-α-1,2-L-Rha) repeating units which are frequently branched with side chains of branched arabinan, β-1,4-galactan or arabinogalactan at the C-4 position. Pectins are synthesized in Golgi apparatus by at least 53 different Golgi-localized glycosyl-transferases (GTs), which transfer glycosyl residues from nucleotide-sugars onto polysaccharide acceptors, and then being secreted into the cell walls. Some pectic glycosyl residues are modified by methyltransferase-catalyzed esterfification or acetyltransferase-catalyzed acetylation during the biosynthesis in the Golgi apparatus (Mohnen, 2008).
Pectin methylesterases (PMEs)
PMEs are ubiquitous enzymes which catalyze the specific hydrolysis of the methyl ester bond at C6 of a GalA residue in the linear HG domain of pectin and releasing methanol and protons. PME (E.C. 3.1.1.11) belongs to CAZy class 8 of carbohydrate esterases (Cantarel et al., 2009). The enzymes were found in plants, phytopathogenic fungi and bacteria as well as insect and yeasts (Gainvors et al., 1994; Lievens et al., 2002; Ma et al., 1990; Markovič and Janeček, 2004).
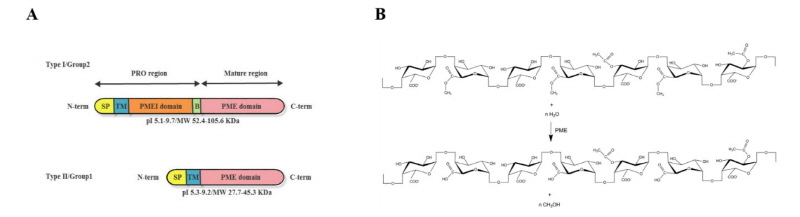
Despite all PME isoforms differ in isoelectric pH (the lowest pI is 3.1 in fungal PME) and the highest pI is 11 in tomato PME), molecular weight and biochemical properties, they catalyse the same reactions of de-methyleste-rification of pectins (Benen et al., 2003). In higher plants, PMEs can be classified into Type I/group2 and Type II/group1 depending on the presence or absence of a N-terminal PRO region which shows the high sequence similarity to PMEI domain of PMEI proteins (Fig. 2A) (Camardella et al., 2000). In detail, Type I/group2 PMEs possess 1-3 PRO domains, referred as long N-terminal PMEI domains, whereas Type II/group1 PMEs contain no PRO domains (PMEI domains). There are no PRO regions found in bacteria and fungi PMEs (Markovič and Janeček, 2004). The PRO regions were supposed to have several hypothetical roles such as supporting in correct conformational folding of the mature PMEs, targeting PMEs to the cell walls, or auto-inhibition PME activity to prevent premature demethylesterification of pectin before secretion into plant cell walls (Micheli, 2001). PMEs extracted from plant cell walls have no PRO region, suggesting that they are cleaved before section, which probably happens inside cells or immediately after export (Dorokhov et al., 2006). The Type I/group2 PMEs have one or two conserved sequences (RRLL, RKLL, KKDL or KRLL, called as Binding Motif for cleavage) severed as cleavage sites recognized by subtilisin-like serine proteases.
Catalytic properties of PME
The substrates of PMEs are methyl-D-galactopyranosyl residues of pectins. PME catalyses the hydrolysis of methyl-ester bonds at C6 of methyl-D-galactopyranosyl residues of pectin, forming negatively charged polymer groups and methanol (Fig. 2B). PMEs may catalyze demethylesterification randomly (fungus PMEs) or linearly (plant PMEs) along the chain of pectins, resulting in opposite consequences on the plant cell walls. If the PMEs act randomly, the demethyesterification leads to random distribution of free carboxyl groups on HGs which are more susceptible to the degradation by plant derived pectinase such as pectate lyases (PLs) and polygalacturonases (PGs) (Kars et al., 2005; Micheli, 2001). The action of pectinase in cooperation with PMEs contributes to cell wall extension and loosening (Moustacas et al., 1991). In contrast, linearly demethylesterification on HGs by PMEs can create long stretches of free carboxyl groups. The blocks of at least 10 consecutive free carboxyl groups will interact with Ca2+, creating a pectic gel (Al-Qsous et al., 2004; Goldberg et al., 1996). In the gel-like environments, the action of endopolygalacturonases is greatly limited, which contributes to cell wall stiffening. Loosening or stiffening of cell walls induced by demethylesterification of PMEs plays important functions in cellular elongation, adhesion, separation and maturation (Jolie et al., 2010; Pelloux et al., 2007). Depending on pH and initial degree of methylesterification of pectins, PMEs can act random or linear demethylesterification, resulting in loosening or stiffening of cell walls. Some PMEs demethylesterifed pectins randomly at acidic pH and linearly at alkaline pH. Moreover, some PMEs show more effective on highly esterified pectin than others at a given pH. The catalytic activity can be enhanced by cations and their concentrations. Trivalent cations catalyze more effectively than bivalent cations , which, in turn, are more effective than monovalent cations (Schmohl et al., 2000). In addition, de-esterified pectins also affect their interaction with other cell wall components such as pectin-cellulose assembly, influencing microfibrils deposition and alignment in plant cell wall (Chanliaud and Gidley, 1999; Yoneda et al., 2010).
Pectin Methylesterase Inhibitor (PMEI)
In addition to regulation by posttranslational mechanism, the PME activity also be regulated by specific proteinaeous inhibitors named PMEI, firstly discovered in kiwi (Actinidia deliciosa), which inhibits against several plant PMEs including PMEs from kiwi, banana, apple, tomato, potato, carrot, apricot and orange. However, they cannot inhibit fungal and bacterial PMEs because the active site clefts in microbial PME structures are deeper than those in plant enzymes. (Balestrieri et al., 1990; D'Avino et al., 2003). The inhibition of PMEIs is fulfilled by forming a PME-PMEI complex in which the PMEI covers the enzyme active site (pectin binding cleft) of PME (Di Matteo et al., 2005). PMEIs are encoded by a large multigene family. There are 71 isoforms in Arabidopsis thaliana, 54 in Poplus trichocarpa, 95 in Linum usitatissimum, 78 in Solanum lycopersicum and 49 in Oryza sativa. The PMEIs transcripts were abundant in different tissues, developmental stages and in response to diverse stress conditions (Nguyen et al., 2016; Pinzon-Latorre and Deyholos, 2014). PMEIs are suggested to have functions in modulating degree of methylesterification on pectins through the inhibitory activity on PMEs. Therefore, PMEIs are probably involved in diverse functions related to PME in plant developments and defenses.
PMEI structure
The three-dimensional structure showed that kiwi PMEI is helical protein, folding in an up-and-down four-helical bundle including α1, α2, α3 and α4 at C-terminus. These helices are aligned in an antiparallel manner and play function as central domain in PMEI structure. Stabilization of the protein structure are strongly supported by hydrophobic interactions at the internal spaces of the helical bundle and disulfide bridge between Cys74 and Cys114, connecting helices α2 and α3. There are three short and distorted helices named αa, αb and αc at the N-terminal region of the PMEI protein structure. Two helices, αa and αb, are connected by disulfide bridge between Cys9 and Cys18. These extended helices are laid at the outside of the central domain and roughly parallel to the plane designated by the helices α1 and α4. The inhibitory on PME activity is obtained by the formation of a reversible, non-covalent 1:1 complex between PME-PMEI in which the inhibitor covers the active site of the enzyme (Giovane et al., 1995; Jolie et al., 2010). In detail, the three helices α2, α3 and α4 interact with the enzyme in proximity of the active site. The N-terminal region of the inhibitor may function in structural stability of the inhibitor. The inhibitor covers the active site cleft of the enzyme preventing the substrate access as well as impeding the interaction for substrate binding. In summary, PMEI prevents PME from binding to the pectin substrate from which inhibits the enzyme activity. Neither PME nor PMEI undergoes dramatic conformational changes when the complex is formed (Camardella et al., 2003; Di Matteo et al., 2005). The lack of inhibition of PMEIs on microbial PMEs activity can be explained. The main reasons are the putative binding site cleft of microbial PMEs is deeper than in plant enzymes and the absence of conserved residues important for interaction with PMEIs in amino acid sequence of microbial PMEs.
PME and PMEI interaction
PME-PMEI interaction is non-covalent and reversible (Balestrieri et al., 1990; Giovane et al., 1995). Neither PME nor PMEI undergoes dramatic conformational changes when the complex is formed (Camardella et al., 2003; Di Matteo et al., 2005). The stability of the complex is strongly impacted by pH, temperature, high-pressure and NaCl concentration. In studies on the complexes of kiwi PMEI and tomato or carrot PMEs, strong interaction was observed at pH <7 and at low salt concentrations (Jolie et al., 2010). High temperature triggers the denaturation of the PME-PMEI complex whereas elevated pressure dissociates the complex from which inactivates the enzyme and inhibitor gradually (Mei et al., 2015). For instance, The inhibitory activity of rice PMEIs against orange PME was reduced when the temperatures were elevated (Nguyen et al., 2016). The carrot PME-kiwi PMEI complex formed large particles aggregating as a single entity during heat treatment (55-65°C), whereas high pressure treatment led to disassociated the complex prior to denaturation of PME alone (Jolie et al., 2010). The studies of the factors involved in controlling the PME-PMEI interaction are very important for applications in plant-based food industries such as cloud stability, textures, viscosity of products. In addition, the specific PME-PMEI pairs were also studied in vivo in plant physiological researches as precious data for understanding the fine-tuning of pectin methylesterification resulted in cell wall modification in plant developments and defense responses. Due to gene abundances and specific inhibition of plant PMEs by plant PMEIs, the studies for specific PME-PMEI pairs are mostly based on co-expression analysis. Until now, there some potential PME-PMEI pairs were reported, including (i) AtPPME1(At1g69940)-AtPMEI2(At3g17220) in Arabidopsis pollen tube (Röckel et al., 2008), (ii) AtPME3-AtPMEI7 interaction in Arabidopsis dark-grown hypocotyls (Sénéchal et al., 2015b), (iii) the possible interaction of AtPME17 with AtPMEI4 functions in root development (Sénéchal et al., 2015a), and (iv) SolyPME1(P14280)-SolyPMEI in tomato fruit ripening (Reca et al., 2012).
Functions of PMEs and PMEIs in plant development
PMEIs are involved in diverse roles in plant physiology. The basics of these involvements are pectin modifications (degree and pattern of HG methylesterification) from which change several cell wall properties (such as elasticity, hydration, porosity and adhesion). Functions of PMEs and PMEIs in plant development, morphology and stress responses will be discussed in detail. Plant development and morphology are strongly influenced by several cell wall properties, including elasticity, porosity, hydration and adhesion, which are strongly linked with pectin composition and modification in an individual cell or an entire tissue (Levesque-Tremblay et al., 2015a; Palin and Geitmann, 2012). Pectin has been demonstrated to be involved in diverse plant physiological processes, including development, growth, differentiation, and defense responses (plant morphogenesis, organogenesis, growth, development, cell-cell adhesion, defense, leaf abscission, fruit maturation and dehiscence, seed hydration, and ion binding) (Mohnen, 2008). It is though that pectin loosening is permissive of growth and hydration, while pectin stiffening restricts growth, required for strength and rigidity (Levesque-Tremblay et al., 2015a). The principle for these functions mostly replies on the hydration and stiffness of pectin matrix, resulting from the methylesterication modification of HGs by the action of PMEs along with the inhibitors, PMEIs. PMEs catalyze the removal of methyl group on HG residues, releasing free carboxyl group on pectins and methanol emission, resulting in the degree of methylesterifcation (DM) modulation. Elevated PME acidity accompanied with low DM pectins. Depending on the pattern of methylesterification (PM) on pectins and the mode of action of PMEs, the de-methylesterification action may result in contrary consequences in cell wall properties. The acidic PMEs randomly de-esterified on HGs, releasing protons which promote action of cell-wall-degrading enzymes (CWDEs) such as endopolygalacturonases (PGs), resulting in cell wall loosening. And, the alkaline PMEs act linearly on HGs, producing consecutive free carboxyl groups with which the Ca2+ interact to create a pectic gels (egg-box formation). The action of CWDEs is limited in gel environment, thus, the action of alkaline PMEs contribute to cell wall stiffening. In addition, PME activity also be regulated by hormones (Micheli, 2001). The inhibitory activity of PMEIs on specific PMEs results in reverse effects on cell wall properties. Therefore, cell wall loosening or stiffening related to methylesterification of pectins are regulated depending on six factors: (i) cells or tissues types, deciding the methylesterification state of substrate in which methylesterified groups on HGs contributed randomly or continuously; (ii) different mode of action of respective PMEs; (iii) the specific inhibitory on PME activities by PME-PMEI complex; (iv) environmental conditions, change in local pH can make alkaline PMEs act randomly on HGs or pectin gel forming in the present of Ca2+; (v) regulations by hormones and (vi) transcriptional regulations, various PME and PMEI isoforms showed differential expressions, both temporarily or partially. The simple action of PME, demethylesterification, involved in diverse plant developmental processes based on these regulations, and the direct or indirect involvements of PMEs and PMEIs in different stages of plant development are explained in the following parts.
Seed germination
Generally, germination starts when seeds imbibe water in an appropriate temperature and receive germination signals in dormant seeds; the process will end when the radicle protruding out from covering layers (Bewley, 1997). Following the water absorption, the seeds activate enzymes in embryos to digest reserve substances in endosperms, supplying energies for embryo growths, resulting in germination. In endospermic species, cell wall remodeling is an essential requirement for seed coat rupture followed by endosperm rupture, completing germination process (Manz et al., 2005; Müller et al., 2006, 2013). In addition, once the seed hydrates, the seed coat will release hydrophilic mucilage composed primarily of pectin. The mucilage was synthesized in Golgi apparatus and secreted into the apoplastic space. During germination process, PME activity is tightly controlled. Altered PME activity will affect the seed germination. The rising evidences of methylesterification of pectin in seed germination will be discussed below. Overexpression of PMEI5 (Ox-PMEI5) showed decreased levels in PME activity in seeds during germination. The Ox-PMEI5 seeds germinated, including testa rupture as well as endosperm rupture, faster than WT seeds did. Low PME activity helps seeds less sensitive to inhibitory of ABA on germination (Müller et al., 2006), suggesting high PME activity along with ABA function in delay of seed germination. The expression of PME and PMEI is temporarily and spatially (tissue-specific manner) regulated during seed germination of garden cress (Lepidium sativum) (Scheler et al., 2015). In addition, PME activity in Chamaecyparis nootkatensis was detected in seeds undergoing to dormancy breakage and germination instead of dormant seeds. Thus, PME activity was positively correlated to the degree of dormancy breakage of yellow cedar seeds. PME was hypothesized playing a critical role in weakening of the mega-gametophyte, allowing radicle emergence and the completion of germination (Ren and Kermode, 2000). Therefore, PME activity is involved in controlling seed germination in both angiosperms and gymnosperm.
Seed mucilage, mainly composed of pectin, is tightly controlled by de-methylesterification on its release. In the seed coat of Arabidopsis, specific expression of PME58 was detected in mucilage secretory cells at the early stages of seed development. The pme58 mutants showed the increased degree of methylesterification HGs due to decrease in PME activity, resulting in decreased mucilage secretory cells surface, and adherent mucilage was less stained by ruthenium red when compared to wild-type seeds, but only in the presence of EDTA, a Ca2+ chelator, suggesting PME58 activated the loosening of the mucilage (Turbant et al., 2016). Another study of the inhibitor of PME, PMEI6, showed the same control manner on mucilage release. Enzyme activity assay showed that PMEI6 inhibited endogenous PME activities, resulting in reduced overall methylesterification of mucilage fractions and in demucilaged seeds (Saez-Aguayo et al., 2013). Taken together, PMEs appear to play a positive roles in the seed mucilage release, while PMEIs inhibit the process.
Organ formation in shoot apical meristem (SAM)
The shoot apical meristem (SAM), located at the tip of the shoot axis, contains a population of cells with characteristics of stem cells dividing to constantly replenish the meristem and to provide cells which will differentiate into plant organs and tissues. Unlike animal stem cells, divided cells from SAM have the unlimited potential in differentiation, such as producing lateral organs, stem tissues or regenerates itself. Specially, SAM mainly controls the pattern of lateral organs initiation, called phyllotaxis. Peaucelle et al. reported that the regulation of de-methylesterification of pectin is a key event in phyllotaxis at SAM. Overexpression of PMEI3 (At5g20740) in the SAM prevented lateral organ formation, whereas overexpression of PME5 (At5g47500) resulted in increased number of primordial. In addition, ectopic overexpression of AtPME5 resulted in abnormal phyllotaxis and in an increase of the number of primordial (Peaucelle et al., 2008). The PME5 activity could be controlled by transcription factor BELLRINGER (BLR) which established normal phyllotaxis (Peaucelle et al., 2011). Thus, regulation of pectin de-methylesterification is necessary and sufficient for organ formations in inflorescence meristems. The meristem is thus another example of a system in which a low degree of pectin methylesterification status leads to a decrease in cell wall stiffness.
Wood development
In dicots, plant growth is not only supported by primary growth but also though secondary growth, which makes plant thicker and wider. Secondary growth is the outward or lateral growth of plants which need an extra support to grow taller than others. Xylogenesis or the formation of wood is typical secondary growth, which produces strongly support for tall plants and wood for human utilization. Wood, secondary xylem, has been commercially exploited as an essential raw material (timber, paper or pulp) or a renewable source of fuel and energy. Secondary xylem is formed by the vascular cambium, a thin layer of stem cells forming secondary xylem and phloem. Wood cells grow in diameter by a combination of intrusive and symplastic growth. There are three requirements for intrusive tip growth requires: dissolution of the middle lamella, yielding of the wall between adjacent cells to create space for the growing tip, and wall biosynthesis at the fiber tip. In wood development, PMEs regulates the fiber width and length in wood of aspen trees due to the demethylesterification activity in middle lamella (Siedlecka et al., 2008). Co-localization of de-methylesterifid pectin and calcium in the middle lamella results in stiffening, on the contrary, the localization of de-methylesterifid pectin is not correlated with calcium, facilitating cell growth. PME genes have been suggested that they play an important role in the development of xylem and cambial cells (Micheli et al., 2000). The role of pectin modification regulated by PME activity in wood development has been suggested when a number of PME isoforms with differential expression pattern and functions was detected at cambial zone in stems of hybrid aspen (Micheli et al., 2000). The abundance of PMEs acts as a negative regulator of both symplastic and intrusive growth of developing wood cells. A study on transgenic modification of PttPME1 expression showed that the activity of PttPME1 inhibits both symplastic growth and intrusive growth by generating methyl-free HG stretches from which creates stiff pectin-calcium-pectin structures and stabilizes pectin network. Thus, PttPME1 would strengthen cellular adhesion and hinder intrusive growth, suggesting that PttPME1 is involved in mechanisms determining fiber width and length in the wood of aspen trees (Siedlecka et al., 2008). In addition, PttPME1 was also involved in tension wood (TW) formation in Populus tremula due to its specitic expression in cambial region of TW (Andersson‐Gunnerås et al., 2006).
Pollen formation and germination
Pollen formation is started by the division of an initial cell, developing into a pollen mother cell (PMC) whose wall composes of primary cell wall, constituting from cellulose, hemicellulose and pectin, and secondary cell wall, entirely made by callose (Francis et al., 2006). PMC undergoes meiotic division, resulting in four microspores which develop within the surrounding callose layers, and then, are warped in an additional wall layers called the primexines. Once the microspores mature, the surrounding walls, including primary and secondary cell walls, are degraded, releasing individual microspores into anther locules within which microspore primexines mature to form the exines, and the outermost pollen walls encapsulate the developing pollen grains. Finally, the pollen walls are completely developed by forming intine walls, inner cellulose walls of pollens, which are located between the exines and the plasma membranes (Boavida et al., 2005; Owen and Makaroff, 1995). The failure in microspore separation during late stages of pollen development was observed in mutations in the QUARTET (QRT) loci in Arabidopsis, resulting in pollen grains released in tetrads (Preuss et al., 1994; Rhee et al., 2003). Among these QRTs, QRT1 encodes a PME, functioning upstream of QRT3 which expresses a polygalacturonase (PG) ((Francis et al., 2006; Rhee et al., 2003) in PMC primary wall degradation. QRT1 optimally catalyzed de-methy-lesterification at slightly acidic pH of 6.3, suggesting the products are randomly de-methylesteried HGs which are the substrates for PG, resulting in loosening the cell wall. These evidences highlight the role of PME in pollen development. However, the promoter analysis of QRT1 showed that the gene was active in a variety of developmentally unrelated tissues, including developing guard cells, the hypocotyl-root transition zone, areas of lateral root emergence and floral nectaries implying the diverse roles of PME activity in plant physiology (Francis et al., 2006).
After pollination by insects, wind or other means, pollen grains, containing the male gametes, are landed on the female stigma. Like seeds, pollen grains are severely hydrated and germinate following the landing on the stigmatic tissues if the pollen-stigma interaction is compatible. During the germination, they grow through the stigma via fast growing pollen tubes in which the male gametes travel downward to fertilize the ovules, resulting in seed forming. PME48, a male gametophyte- specific PME, impacts the mechanical properties of the intine wall during maturation of the pollen grain, in turn, influencing pollen grain germination (Leroux et al., 2015). Pollen grains of homozygous PME48 mutant showed two tips emerging instead of one in the wild-type, and displayed a significant delay in imbibition and germination. The mutant decreased in PME activity accompanied with increase degree of methyltesterification HGs. The Ca2+ supplement can restore the wild-type phenotype on PME48 mutant pollens, suggesting de-methylesterified HGs is a Ca2+ source which necessary for normal pollen germination. Finally, Leroux et al. suggested the model showing the involvement of PME48 in pollen dehydration, imbibition and germination. During the maturation of pollen grains, PME48 may be secreted into the intine wall and catalyze the methylesterified HGs by random or block-wise actions, resulting in weakly de-methylesterified HGs which could form pectate-gel with Ca2+. Fowling the landing on stigma tissues, the pollens will uptake water, called imbibition of the pollen grains, enhanced by the presence of the more hydrophilic weakly de-methylesterified HGs. And then, calcium influx from the partial pollen cell wall to pollen grains release of Ca2+ from the pectate gel, thus weakening the mechanical properties of the intine wall which may be degraed by PGs or PLs and/or break under the turgor pressure. Therefore, these processes may promote the emergence of the pollen tube tips, resulting pollen germinations (Leroux et al., 2015). Pollen tube elongation occurs in self-similar manner in which the hemisphere tip maintains its shape, while expanding and producing a constant diameter cylinder at its base due to the tightly control in action of PMEs and PMEIs on the methylesterfied pectin. The tip growth is induced by the deposition of methylesterified pectin and the present of PMEIs, inhibiting the de-methylesterification of pectin, thus, resulting in tip growth. And, at the cylindrical base and the shoulder of the apical dome, the action of PMEs along with the present of Ca2+ trigger the pectin-gel formation by cross-linking between de-methylesterified pectin and Ca2+, preventing any further cell wall expansion. Any alterations within the processes of pollen tube elongation will resulted in negative defects in the pollen growth. For instance, excess applying of exogenous PME arrests pollen tube growth due to thickening of the apical cell wall (Bosch et al., 2005; Parre and Geitmann, 2005). Interestingly, mutation of a pollen PME isoform in Arabidopsis, VANGUARD1, reduced PME activity in the pollen, greatly retarded pollen tube growth in style and transmitting tract, abolished the penetration to the female tissue, thus, reducing male fertility (Jiang et al., 2005). Interference with other pollen PMEs showed similar effects. In particularly, the lack of Arabidopsis pollen-specific PME, AtPPME1 (Atg69940), expression resulted in reduction of the overall PME activity in pollen grain of the atppme1 plants, and in vitro germination test showed that Wild-type pollen tubes grew straight, while pollen tubes of the atppme1 mutant exhibited curved, irregular morphology, dramatically stunted and remained shorter than the wild-type (Tian et al., 2006). In addition, PMEI isoform, ATPMEI2 specifically expressed in Arabidopsis pollen and physically interacted with AtPPME1, locally inhibiting PME activity and regulating cell wall stability at the pollen tube tip (Röckel et al., 2008). Moreover, the silencing of NtPPME1, a pollen specific PME from Nicotiana tabacum, resulted in a mild but significant decrease of in vivo pollen tube growth while the overall PME activity in pollen was not significantly affected (Bosch and Hepler, 2006). Taken together, the PME activity is restricted controlled at the tip of the pollen tube, together with the addition of methylesterifed pectin facilitated cell wall expansion for tip growth. Whereas, at the shoulder and the trunk of the tip, the PME activity is strongly dynamic action in de-methylesterification of pectin, resulting pectin-calcium formation, strengthening the cell wall at the base of pollen tube. These events are critical for the pollen tube growth. In summary, the activities of PMEs localized at the pollen tube’s apex are specifically inhibited by PMEIs, resulting higher DM pectin, therefore are conducive to pollen tube elongation.
Fruit development and ripening
Developmental stages of fruit include early stages of fruit development, 0-35 day-after-ferilization (DAF), and ripening stages, mature green (MG), breaker (Br), turing (Tu), ripe (Ri) and overripe (RR). Different PME isoforms are expressed during tomato fruit development, such as two PME isoforms (PME1 and PME2) are expressed in both mature green and ripe fruits. However, PME1 decreases slightly in the ripen fruits, whereas PME2 increases roughly two-fold from the MG stage to the Ri stage (Tucker et al., 1982). PME expression and activity in tomato pericarp is highly regulated during fruit development (Harriman et al., 1991). The level of PME activity was lowest at 10 (DAF) fruit, and highest at the turning stage of fruit development. During early stages of fruit development, the increase in PME activity parallels to the increase in PME protein. However, between the Tu stage and Ri stage, the PME activity was decreased, while the PME protein was still increased. PME protein was first detected in 20-d-old fruit and levels of PME protein increased gradually until the ripe stage. At the late stage of fruit ripening, the PME activity was reduced. The decrease in PME's specific activity indicates another level of control, such as PME inhibitors. For instance, SolyPMEI, a PMEI in tomato, specific interacts with PME1, inhibiting PME activity and playing a role in the spatial distribution of esterified pectin in the fruit (Reca et al., 2012). In addition, silencing of PMEU1, a ubiquitous PME, in tomato showed faster fruit softening during the ripening, suggesting the PMEU1 functioned in strengthen the cell wall (Phan et al., 2007). Similar to tomato, the specific expression of MaPMEI gene in banana fruits indicates its possible role in the regulations of PMEs (Srivastava et al., 2012). Moreover, there is a significant decrease in tissue integrity during fruit senescence of ripen fruits from transgenic tomato plants with reduced PME activity, and the fruits are more susceptible to microbial attack (Tieman and Handa, 1994), suggesting the function of PME on tissue integrity.
Conclusion
Pectin methylesterification status, controlled by PME activity and its inhibitor named PMEIs, clearly impacts diverse plant developmetal processes and stress responses. PMEs play major roles in modification of pectin properties, such as the stiffening by forming Ca2+-pectate cross-link complexes or loosening by triggering CWDEs to break out cell wall components including pectin and its interaction with other cell wall components. By inhibition of PME activity, the inhibitor PMEIs act in opposite consequences on pectin properties, leading to contrary phenotypes between the action of each individual in PME-PMEI pairs. Despite the importance of the PMEs and PMEIs in plant development, environmental responses and biotechnology applications, there have been only a few genes of these large families characterized and utilized. Therefore, studying on PME and PMEI families in all plant species is necessary to discover more physiological functional PMEs or PMEIs. Gene redundancy and the complicated regulation mechanisms in these families are either the big challenge or interesting missions for biologists who are eager to discover the mysteries of plant biology.