Introduction
Materials and Methods
Isolation, Cultivation, and 16S rRNA Gene Analysis
Genome Sequencing and Functional Annotation
Phenotypic and Biochemical Characterization
Chemotaxonomic Analysis
Results and Discussion
Phylogenetic Position and Genomic Features
Plant Growth-Promoting Traits
Biochemical and physiological characterization
Safety Assessment
Physiological Characteristics
Conclusion
Introduction
Sustainable crop production increasingly depends on environmentally sound alternatives to chemical fertilizers and pesticides. Plant growth-promoting bacteria (PGPB) are promising biological resources that enhance nutrient availability, stimulate root development, and improve plant tolerance to abiotic stresses. Among them, Enterobacter species have been reported to contribute to nitrogen fixation, phosphate solubilization, and phytohormone synthesis, including indole-3-acetic acid (IAA), which collectively promote plant growth under diverse environmental conditions (Andrés-Barrao et al., 2017; Mezzatesta et al., 2012). However, because several Enterobacter species are also known opportunistic pathogens, their application as agricultural inoculants requires careful genomic and phenotypic safety assessment (Doijad et al., 2016).
In recent years, insect-associated microbiomes have emerged as valuable reservoirs for discovering metabolically versatile and stress-tolerant microbes. The gut environments of wood-feeding insects, in particular, impose strong selective pressures, including nutritional limitation, high lignocellulosic content, and exposure to toxic plant secondary metabolites. These constraints frequently enrich for bacteria capable of decomposing recalcitrant biomass and synthesizing essential nutrients (Ceja-Navarro et al., 2019; Engel and Moran, 2013; Yun et al., 2014). Monochamus alternatus, the major vector of pine wilt disease, harbors a diverse gut microbiota dominated by Gammaproteobacteria, including Enterobacter species, which support host nutrition during larval feeding on pine xylem (Ge et al., 2020).
Given these ecological adaptations, insect gut bacteria represent promising candidates for identifying PGPB with robust physiological resilience suited for fluctuating soil environments. In this study, we isolated the bacterial strain MALB-1 from the gut of M. alternatus larvae and performed a polyphasic characterization to determine its taxonomic position and functional attributes. Although MALB-1 exhibited high 16S rRNA gene similarity to known Enterobacter species, whole-genome analyses, including average nucleotide identity (ANI) and digital DNA-DNA hybridization (dDDH), revealed clear genomic distinctiveness, warranting further taxonomic, biochemical, and chemotaxonomic evaluation (Lee et al., 2016; Meier-Kolthoff et al., 2013).
Beyond classification, we comprehensively examined MALB-1 for traits associated with plant growth promotion—such as nitrogen metabolism, phosphate solubilization pathways, and IAA-related genes—and evaluated its physiological tolerance to agriculturally relevant stress conditions. Additionally, recognizing biosafety concerns associated with the genus Enterobacter, we screened for virulence factors and antibiotic resistance genes using established genomic databases (Chen et al., 2005; McArthur et al., 2013) and performed phenotypic antibiotic susceptibility analyses. Through this integrative approach, we aimed to assess whether MALB-1 possesses the combined attributes of functional efficacy and biosafety required for development as a practical agricultural inoculant.
Materials and Methods
Isolation, Cultivation, and 16S rRNA Gene Analysis
Strain MALB-1 was isolated from the gut of M. alternatus obtained from OsangKinsect Co., Ltd. (Namyangju, Korea). The isolated strain was routinely cultured on Luria-Bertani (LB) agar (MBcell, Seoul, Korea) at 30°C for 48 h. The 16S rRNA gene sequence was obtained using PCR with universal primers 27F and 1492R synthesized by Macrogen Inc. (Seoul, Korea) (Lane, 1991). The sequence was aligned using CLUSTAL_W (Thompson et al., 1994), and a phylogenetic tree was constructed using the maximum-likelihood method (Felsenstein, 1981) in MEGA-X (Kumar et al., 2018). Evolutionary distances were computed using the Kimura 2-parameter model with a discrete Gamma distribution, and the robustness of the tree topology was evaluated using bootstrap analysis with 1,000 replications (Kimura, 1980).
Genome Sequencing and Functional Annotation
Whole genome sequencing was performed using the PacBio Sequel II platform (Pacific Biosciences, Menlo Park, CA, USA). The assembled genome was annotated to identify functional genes, and plant growth-promoting genes were screened based on Clusters of Orthologous Groups (COG) functional categories. For taxonomic confirmation, Average Nucleotide Identity (ANI) values were calculated using OrthoANI (Lee et al., 2016), and digital DNA-DNA hybridization (dDDH) values were determined using the Genome-to-Genome Distance Calculator (GGDC 2.1) (Meier-Kolthoff et al., 2013). To assess genomic safety, putative virulence factors were screened against the Virulence Factor Database (VFDB) (Chen et al., 2005), and antibiotic resistance genes were identified using the Comprehensive Antibiotic Resistance Database (CARD) (McArthur et al., 2013).
Phenotypic and Biochemical Characterization
Biochemical characteristics were analyzed according to the manufacturer’s instructions. Enzyme activities and carbon source utilization were tested using API 20E and API 50CH kits (bioMérieux, Marcy-l’Étoile, France) and GEN III Microplates (Biolog, Hayward, CA, USA). Growth characteristics were evaluated at various temperatures (10-50°C), pH levels (pH 4-10), and NaCl concentrations (0-8% w/v) to determine the physiological range. Hydrolytic activity was assessed using substrates including casein (Acumedia, Lansing, MI, USA), Tween 20 (Daejung Chemicals, Siheung, Korea), Tween 60 (Daejung Chemicals, Siheung, Korea), and starch. Antibiotic susceptibility was determined using the disc diffusion method (Bauer et al., 1966). Strain MALB-1 and related reference strains were tested against antibiotics including tetracycline, chloramphenicol, ampicillin, kanamycin, streptomycin, and norfloxacin to compare resistance profiles.
Chemotaxonomic Analysis
For cellular fatty acid analysis, cells were harvested after cultivation on LB agar at 30°C for 48 h. Fatty acids were extracted through saponification and methylation processes and analyzed using gas chromatography according to the Sherlock Microbial Identification System (MIDI Inc., Newark, DE, USA) (Sasser, 1990).
Results and Discussion
Phylogenetic Position and Genomic Features
The 16S rRNA gene sequence analysis showed that strain MALB-1 belongs to the genus Enterobacter, exhibiting the highest similarity to Enterobacter asburiae ATCC 35953T (99.3%) and Enterobacter chengduensis KCTC 52993T (99.3%) (Table 1). In the phylogenetic tree inferred from 16S rRNA gene sequences, strain MALB-1 formed a distinct phyletic lineage separate from other valid Enterobacter species, supported by a high bootstrap value of 100% (Fig. 1). This topological position indicates that MALB-1 forms a distinct phyletic lineage within the genus, suggesting genomic divergence from currently described Enterobacter species.
Table 1
Genomic relatedness indices between Enterobacter sp. MALB-1 and closely related Enterobacter type strains
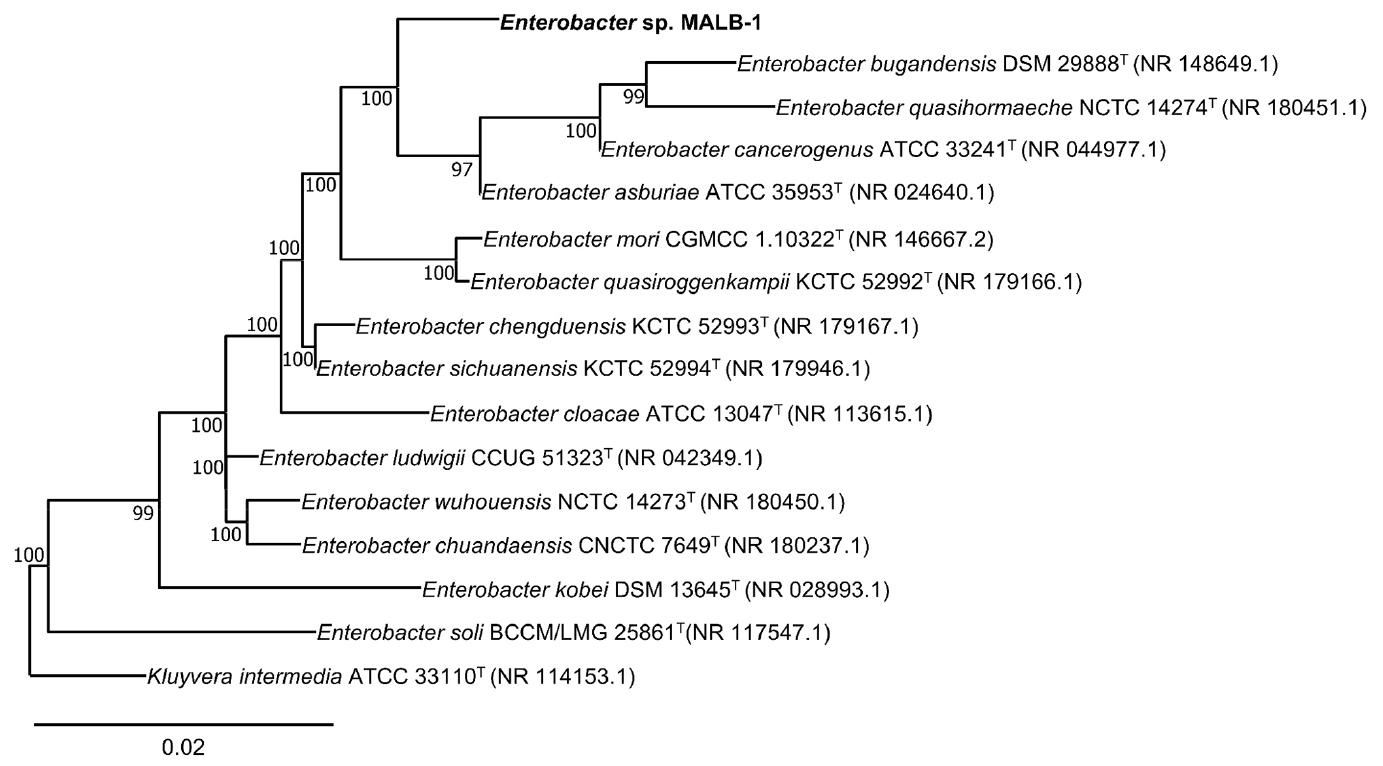
Fig. 1.
Maximum-likelihood phylogenetic tree based on 16S rRNA gene sequences showing the phylogenetic position of Enterobacter sp. MALB-1 relative to other type strains within the genus Enterobacter. The tree was reconstructed using the Kimura 2-parameter model. Bootstrap values (expressed as percentages of 1,000 replications) greater than 70% are shown at the branching points. Kluyvera intermedia ATCC 33110T was used as an outgroup. GenBank accession numbers (PQ060136) are indicated in parentheses. Bar, 0.02 substitutions per nucleotide position.
To clarify the precise taxonomic position, whole-genome relatedness indices were calculated. The Average Nucleotide Identity (ANI) values between MALB-1 and closely related type strains ranged from 86.7% to 88.2%, which is significantly below the generally accepted species demarcation threshold of 95-96% (Lee et al., 2016). Furthermore, the digital DNA-DNA hybridization (dDDH) values were low (31.8-34.7%) (Table 1), unequivocally falling below the 70% threshold required for species definition. Collectively, genomic relatedness indices support that MALB-1 is genetically distinct from currently characterized Enterobacter species, although additional taxonomic analyses would be required for formal species delineation (Parte et al., 2020). Based on these findings, we designated the isolate as Enterobacter sp. MALB-1. The genome size was 4.68 Mb with a G+C content of 54.1% (Table 2).
Table 2
General genomic features of Enterobacter sp. MALB-1
|
Genome size (bp) |
No. of contigs |
DNA G+C content (%) |
Total genes |
Protein coding genes |
RNA genes |
rRNA genes (5S/16S/23S) |
tRNA genes |
| 4.68 Mb | 5 | 54.1 | 4,357 | 4,246 | 111 | 25 (9/8/8) | 86 |
Plant Growth-Promoting Traits
Genomic analysis identified a diverse array of genes associated with plant growth promotion, including those involved in nitrogen metabolism, phosphate solubilization, indole-3-acetic acid (IAA) biosynthesis, and organic acid production (Table 3). These included genes for nitrogen fixation, phosphate solubilization, and carbohydrate metabolism. The presence of genes involved in indole-3-acetic acid (IAA) biosynthesis suggests that MALB-1 can stimulate root growth in plants. Similar PGP traits have been reported in other Enterobacter species, such as Enterobacter sp. SA187, which enhances stress tolerance in plants (Andrés-Barrao et al., 2017).
Table 3
Plant growth-promoting genes identified in the genome of Enterobacter sp. MALB-1
Biochemical and physiological characterization
The metabolic versatility and environmental adaptability of Enterobacter sp. MALB-1 were evaluated using API 20E, API 50CH, and GEN III microplate assays (Table 4). The strain exhibited a broad metabolic profile, capable of utilizing a diverse array of carbon sources. Specifically, MALB-1 effectively metabolized various monosaccharides, disaccharides, and organic acids, which are major components of plant root exudates. This metabolic flexibility suggests that MALB-1 possesses a competitive advantage for colonization and survival in the nutrient-fluctuating rhizosphere environment. In terms of abiotic stress tolerance, the strain demonstrated robust growth under saline conditions (4% NaCl) and acidic environments (pH 5 and 6), indicating its suitability for application in saline or acidic soils. Furthermore, the antibiotic resistance profile revealed that MALB-1 is resistant to specific antibiotics such as vancomycin and lincomycin, a trait that may contribute to its persistence in soil microbiomes containing antibiotic-producing competitors. The detailed biochemical and physiological characteristics are summarized in Table 4.
Table 4
Biochemical and physiological characteristics of Enterobacter sp. MALB-1
(+), Positive / Utilized; (-), Negative / Not utilized; (Res*), Resistant (growth observed in the presence of the antibiotic). Data represent a compilation of results from API 20E, API 50CH, and GEN III microplate assays. Note that citrate utilization was negative in API 20E (24h) but positive in GEN III (extended incubation), indicating potential adaptation or assay-specific conditions.
Safety Assessment
Given the clinical significance of some Enterobacter species (Mezzatesta et al., 2012), a safety assessment is mandatory. The genomic search against the VFDB detected 18 putative virulence genes, including cdtA and lpfC (Table 5). Additionally, 13 putative antibiotic resistance genes were identified via the CARD database (Table 6). These included the tetracycline resistance gene tet(43) and various multidrug efflux pump genes such as marA and ramA (McArthur et al., 2013).
Table 5
Putative virulence factors identified in the genome of Enterobacter sp. MALB-1
VFDB IDs and descriptions are based on the Virulence Factor Database (VFDB; http://www.mgc.ac.cn/VFs/). Genes with a sequence identity of ≥ 60% were included in this analysis.
Table 6
Putative antibiotic resistance genes identified in the genome of Enterobacter sp. MALB-1
ARO, Antibiotic Resistance Ontology; AMR, Antimicrobial Resistance. ARO IDs and descriptions are based on the Comprehensive Antibiotic Resistance Database (CARD; https://card.mcmaster.ca/). Genes with a sequence identity of ≥ 60% were included in this analysis.
Consistent with the genomic predictions, phenotypic characterization confirmed that strain MALB-1 exhibits resistance to tetracycline and chloramphenicol (Table 7). However, notably, the strain demonstrated susceptibility to broad-spectrum antibiotics, including ampicillin, kanamycin, streptomycin, and norfloxacin. This susceptibility profile stands in stark contrast to the comparative Enterobacter strains (ER-2 to ER-5), which displayed extensive multidrug resistance phenotypes across most tested antibiotics. Such a distinction is critical, as it differentiates MALB-1 from hazardous clinical isolates often associated with the genus Enterobacter. Consequently, despite the presence of intrinsic resistance traits, the comparative analysis suggests that MALB-1 poses a significantly lower and manageable risk for agricultural application compared to other members of the genus.
Table 7
Antibiotic susceptibility profiles of Enterobacter sp. MALB-1 and comparative Enterobacter species
Antibiotic susceptibility was evaluated using the paper disc diffusion method. Paper discs (8 mm diameter) were loaded with 12 µL of each antibiotic solution (10 mg/mL) and placed on LB agar plates inoculated with the bacterial strains. The zones of inhibition (halo radius, r) were measured after 48 hours of incubation. Abbreviations: R, Resistant (no clear zone or r < 4 mm); S, Susceptible (clear zone with r ≥ 4 mm). Antibiotics: NOR, Norfloxacin; CEF, Cefotaxime; LEV, Levofloxacin; AMP, Ampicillin; TET, Tetracycline; KAN, Kanamycin; CHL, Chloramphenicol; STR, Streptomycin. Reference Strains: ER-2, Enterobacter quasiroggenkampii; ER-3, Enterobacter aerogenes; ER-4, Enterobacter chengduensis; ER-5, Enterobacter sichuanensis.
Physiological Characteristics
Physiological characterization revealed that strain MALB-1 is capable of growth at temperatures of 10-40°C, pH 5-9, and NaCl concentrations of 0-5% (w/v). Although its tolerance ranges for pH and salinity were slightly narrower or comparable to those of the reference strains, these conditions are well within the typical ranges found in agricultural soils, supporting the strain’s ecological fitness. More importantly, MALB-1 could be clearly distinguished from phylogenetically related Enterobacter species based on specific biochemical traits (Table 8). For instance, MALB-1 was unable to ferment sorbitol and inositol, whereas most reference strains (ER-2, ER-3, and ER-4) utilized these substrates. Additionally, differential traits such as citrate utilization and sucrose fermentation further differentiated MALB-1 from E. quasiroggenkampii (ER-2). Collectively, these differential characteristics, combined with the genomic data, confirm the distinct taxonomic position of MALB-1.
Table 8
Differential characteristics of Enterobacter sp. MALB-1 and related type strains
Strains: ER-2, Enterobacter quasiroggenkampii; ER-3, Enterobacter aerogenes; ER-4, Enterobacter chengduensis; ER-5, Enterobacter sichuanensis. Symbols: +, Positive (growth or utilization); -, Negative (no growth or utilization). Data represent selected differential traits obtained from growth assays, hydrolysis tests, and API 20E systems. Characteristics that were identical across all strains are not shown in this table.
Chemotaxonomic analysis revealed that the major cellular fatty acids of strain MALB-1 were C16:0 (22.1%), Summed feature 3 (C16:1ω7c and/or C16:1ω6c, 24.6%), and Summed feature 8 (C18:1ω7c/C18:1ω6c, 27.7%) (Table 9). This fatty acid profile is consistent with the genus Enterobacter. However, clear quantitative differences were observed compared to the reference strains; for instance, MALB-1 contained a notably higher proportion of C14:0 (5.5%) than the related type strains (0.5-1.5%). These distinctive chemotaxonomic features further support the unique taxonomic status of strain MALB-1.
Table 9
Cellular fatty acid composition (%) of Enterobacter sp. MALB-1 and related type strains
Strains: ER-2, Enterobacter quasiroggenkampii; ER-3, Enterobacter aerogenes; ER-4, Enterobacter chengduensis; ER-5, Enterobacter sichuanensis. Values are percentages of total fatty acids. Major fatty acids (>10%) are indicated in bold. - , Not detected or <0.1%. Summed feature 3 comprises C16:1ω7c and/or C16:1ω6c. Summed feature 8 comprises C18:1ω7c/C18:1ω6c.
Conclusion
This study successfully isolated and comprehensively characterized Enterobacter sp. MALB-1 from the gut ecosystem of M. alternatus, providing empirical evidence that insect-associated microbiomes can serve as a valuable reservoir for agriculturally beneficial bacteria. The primary objective of this research was to define the strain’s taxonomic position using a polyphasic approach, while simultaneously evaluating its plant growth-promoting potential and biosafety for agricultural application.
Taxonomically, our analysis confirmed that MALB-1 represents a unique genomic species within the Enterobacter complex. Despite exhibiting high 16S rRNA gene similarity to E. asburiae ATCC 35953T and E. chengduensis KCTC 52993T, whole-genome sequencing revealed that both Average Nucleotide Identity (ANI) values (86.7-88.2%) and digital DNA-DNA hybridization (dDDH) values (31.8-34.7%) were significantly below the generally accepted thresholds for species delineation. Furthermore, the strain was clearly differentiated from phylogenetically related type strains by its distinct chemotaxonomic profile, characterized by a notably higher proportion of C14:0 fatty acid, and specific biochemical traits, such as the inability to ferment sorbitol and inositol. These findings indicate that MALB-1 represents a genomically and phenotypically distinct Enterobacter lineage. While such distinctiveness implies taxonomic novelty, the primary focus of this study remains on elucidating the strain’s functional potential and ensuring its biosafety for practical application.
From an agronomic perspective, MALB-1 possesses a robust genetic and physiological arsenal suitable for biofertilizer formulation. Genomic analysis identified essential genes for nitrogen fixation, phosphate solubilization, and indole-3-acetic acid (IAA) biosynthesis, which are critical for enhancing soil fertility and crop productivity. Physiological assays further demonstrated that the strain is metabolically versatile and capable of thriving under abiotic stresses, such as saline and acidic conditions. These traits, likely evolved to facilitate survival in the harsh insect gut environment, suggest that MALB-1 is well-equipped to colonize the rhizosphere and compete effectively in diverse soil ecosystems.
Crucially, this study addresses the safety concerns often associated with the genus Enterobacter. Although genomic screening identified putative virulence factors and antibiotic resistance genes, phenotypic assays confirmed that MALB-1 lacks the functional multidrug resistance typically observed in clinical isolates. Unlike the comparative reference strains which exhibited extensive resistance, MALB-1 remained susceptible to broad-spectrum antibiotics including ampicillin and kanamycin. This discordance between genotype and phenotype implies that the identified resistance genes may be silent or expressed at low levels, suggesting that the risk of introducing MALB-1 into agricultural soils is low and manageable.
Collectively, Enterobacter sp. MALB-1 stands out as a highly functional and relatively safe bio-inoculant candidate. To translate these laboratory findings into practical solutions, future research should focus on large-scale in vivo field trials to validate its efficacy and monitor its interaction with indigenous soil microbial communities.




