Introduction
Materials and Methods
Plant materials and cirsimaritin extraction
Cellular RNA extraction and transcriptome sequencing
De novo assembly
Functional annotations
Isolation of cirsimaritin pathway genes
Phylogenetic analysis
Results
Cirsimaritin content
RNAseq results
Functional annotation
GO and KEGG analyses
Isolation of genes in the cirsimaritin biosynthesis
Phylogenetic analysis
Discussion
Introduction
The genus Cirsium is one of the largest genera in Asteraceae. It contains about 250 perennial and biennial species that distribute worldwide (Kadereit, 2007, Yildiz et al., 2016). Some of the Circium species are known as thistles which are common name for a group of flowering plants characterized by having prickles on the margins of leaves and Cirsium species differ from other thistle genera (Carduus, Onopordum, Silybum, Cynara etc) by having feathered hairs on their achenes (Rose, 1981). Ten Cirsium species have been identified in wild in Korea (Song and Kim, 2007). Cirsium pendulum Fisch, known as Korean thistle, is a Korean endemic Cirsium species. Antioxidant activities were reported from various parts of C. pendulum (Chon et al., 2006). Cirsium setidens Nakai is also an endemic Cirsium species and known as its common name Gondrae in Korea where young leaves of Gondrae have long been used for side dish vegetable. C. setidens has herbal medicinal effect such as recovery from fatty liver injury (Kim and Chung, 2016), antioxidant activities (Lee et al., 2016), and neuroprotective (Chung et al., 2016). Thao et al. (2011) analyzed the bioactive flavonoids from selected Korean thistles including C. pendulum Fisch and C. setidens Nakai. Their results revealed six flavonoids luteolin 5-O-glucoside, luteolin 7-O-glucoside, hispidulin 7-O-neohesperidoside, luteolin, pectolinarin, and apigenin and suggested these flavonoids as the chemical markers of the thistles. Cirsium japonicum Maxim is a wild thistle plant in east Asia. It has been widely used for herbal medicine in east Asia including Korea, China, and Japan (Luo et al., 2021). Extracts from C. japonicum Maxim have been known to be effective in treatments of diabetes and Alzheimer’s disease (Wagle et al., 2019), breast cancer (Kim et al., 2010; Park et al., 2017) and other chronic diseases (Mahmood and Alkhathlan, 2019).
Plant secondary metabolites have been highly utilized in pharmaceutical industry for medicinal purposes. They are not essential for plant growth, but plants produce them for coping with biotic and abiotic stresses (Bourgaud et al., 2001). Plant secondary metabolites are categorized into three classes by their structures: terpenoids and steroids, phenolic compounds, and alkaloids (Bourgaud et al., 2001; Hussein and El-Anssary, 2017). Cirsimaritin (4’,5-Dihydroxy-6,7-dimethoxyflavone) is a member of flavonoids in a class of polyphenolic secondary metabolites. It was also known as 7-O-methylated flavonoid in a class of polyphenolic secondary metabolites. Cirsimaritin is a major alkaloid in genus Cirsium (Benali et al., 2022; Lee et al., 2017), but it was also found in other plant species (Mahmood and Alkhathlan, 2019). In the analysis of flavonoid contents in C. japonicum var. Maakii, the compounds in the EtOAc (ethyl acetate) fraction were identified as the cirsimaritin, hisidulin, and cirsimarin, in which the cirsimaritin was the main constituent (Lee et al., 2017).
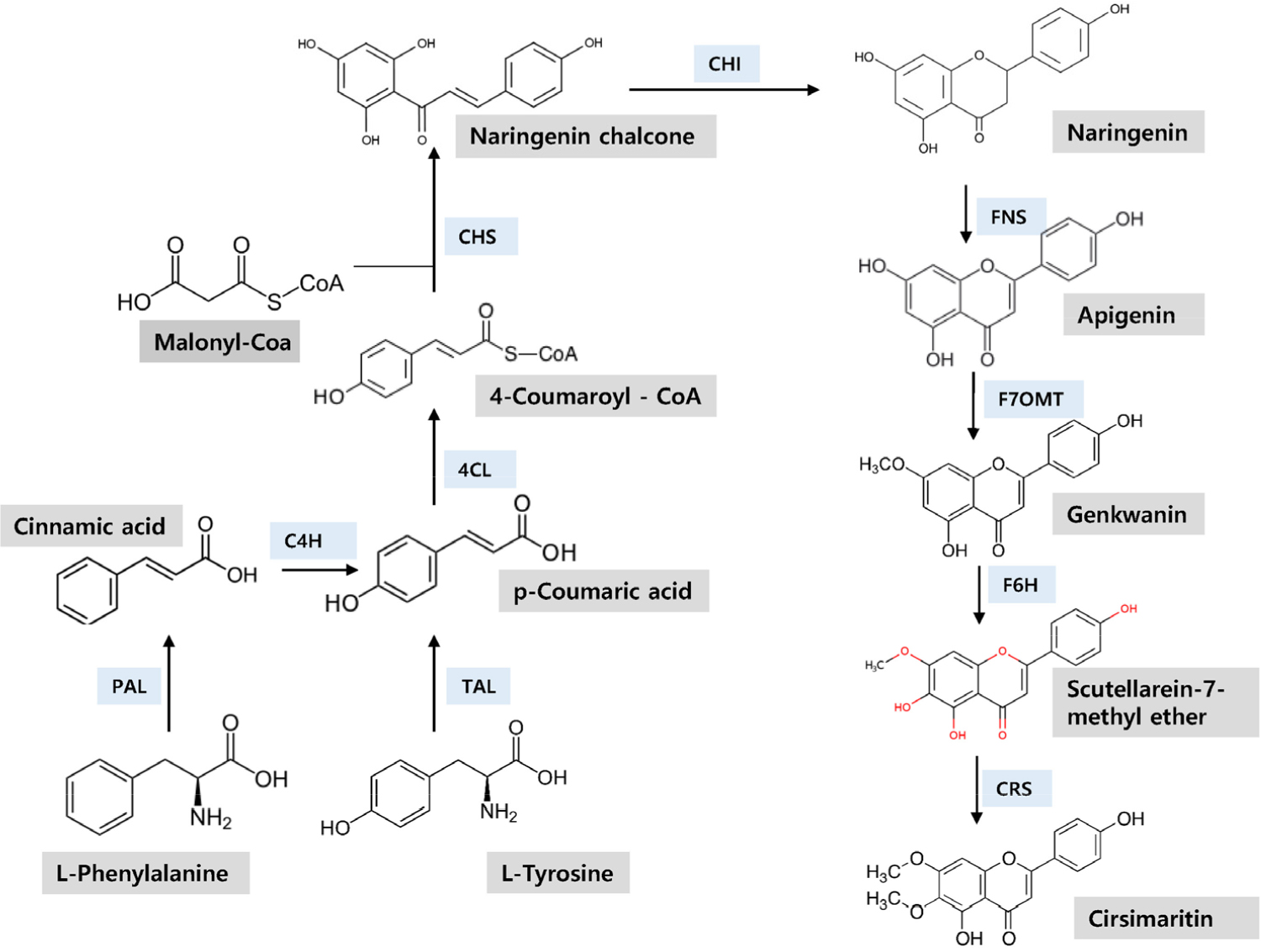
Like many other alkaloid biosynthesis, cirsimaritin biosynthesis starts with tyrosine and also with phenylalanine (Lichman, 2021; Winkel-Shirley, 2001). L-tyrosine convert to P-coumaric acid by tyrosine ammonia lyase (TAL), then, the P-coumaric acid is transformed to P-coumaryl-CoA which is converted to naringenin chalcone by chalcone synthase (CHS) (Kreuzaler and Hahlbrock, 1972). A consecutive enzyme chalcone isomerase (CHI) converts the narignenin chalcone to a flavonoid naringenin by flavone synthase (FNS). The flavonoid naringenin is, then, converted to other flavonoids, apigenin, genkwanin, scutellarein-7-methyl and finally to cirsimaritin by stepwise enzyme meditated reactions. The biochemical synthetic steps from tyrosine to genkwanin were demonstrated with Escherichia coli system by incorporating involved genes (Lee et al., 2015). Of the enzymes involved in the pathway, CHS and CHI are key enzymes for syntheses of chalcone and flavones.
In the current study, we identified all genes in the biosynthetic pathway of cirsimaritin from the transcriptomes from Korean endemic C. setidum Fisch. We present here the molecular detail of the gene involved in the cirsimaritin biosynthetic pathway in Cirsium species. Phylogenetic analyses of the CHS and CHI are also presented. Prior to our study, cirsimaritin biosynthesis genes were reported from a study of transcriptomes of C. japonicum (Park et al., 2020)
Materials and Methods
Plant materials and cirsimaritin extraction
Cirsium plants were obtained from local native plants in Gyeonggi province, South Korea. We extracted cirsimaritin from fresh and dried tissues that were collected when the flowers were full in bloom in middle August 2020. For dried tissue preparation, the tissues were left in a drying oven at 60°C for 48 hours. Then, the tissues were ground with a mortar and pestle. Extraction and measurement of the cirsimaritin were followed by the protocol of Lee et al. (2017).
Cellular RNA extraction and transcriptome sequencing
Cellular RNAs were extracted from fresh tissues of leaf of C. setidum Nakai when the flowers were full bloomed using Hybrid-RTM kit (Genes All Biotechnology Co., Seoul, Korea). cDNA libraries were constructed in each tissue separately using the TruSeq RNA Sample Prep Kit v2 (Illumina) and paired-end sequencing was carried out with the Illumina HiSeq 4000 at Macrogen, Inc. (Seoul, South Korea). We discarded the low quality reads (Phred score < 20), shirt reads (< 50 bp), and empty nucleotides (N at the endof reads). After removed the adapter sequences with Trimmomatic software, sequencing quality was checked using FastQC software (https://www.bioinformatics.babra ham.ac.uk/projects/fastqc/).
De novo assembly
Transcriptomes from three libraries were pooled for de novo assembly with RNA-Seq Assembly Pipeline using Trinity v2.13.2 with the default option (https://github.com/trinityrnaseq/trinityrnaseq/wiki). Duplicated contigs were removed using CD-HIT-EST software (http://weizhong-lab.ucsd.edu/cd-hit/). The low coverage contigs (<10 hits) were removed using Samtools v1.13 (http://www.htslib.org/) with the default option. The coding region was, then, finally confirmed using TransDecoder (v3.0.1) (https://github.com/TransDecoder/TransDecoder/wiki).
Functional annotations
The transcriptome sequences were blasted on the known public protein databases InterProScan in the European Bioinformatics Institute (EBI) and NR in NCBI using the Basic Local Alignment Search Tool (BLAST) program with the cut-off parameters e-value 1e-4 and > 70% similarity. The candidate transcripts with InterProScan and BLASTP hits were sorted to import to the Blast2GO suite 6.0 (https://www.blast2go.com/, BioBam Bioinformatics SL, Valencia, Spain) for gene ontology (GO) analysis which was carried out with an E-value cut-off of 1 × 10-5.
Isolation of cirsimaritin pathway genes
Cirsimaritin biosynthetic pathway was elucidated in Dracocephalum kotschyi (Poursalavati et al., 2021), which were used as query sequences in BLAST analysis in the NCBI database. We downloaded the whole set of protein sequences of each enzyme in the biosynthetic pathway in Arabidopsis thaliana and Glycine max from NCBI, then, selected the representative sequences in each set of enzymes using the option of “Identical protein groups” in the NCBI. The selected set of enzymes sequences were used for query in the transcriptome sequences of Cirsium species using NBLAST program with a selection criteria > 60% similarity in > 100 amino acids (Choi et al., 2022). The genes isolated were blasted again in NCBI database to be sure their functions.
Phylogenetic analysis
The protein sequences were aligned using ClustalW software and the phylogenetic tree was constructed with the neighbor-joining method using MEGA v5.0 with 1000 bootstraps. A maximum likelihood phylogenetic tree was built using MEGA X (v10.2.4).
Results
Cirsimaritin content
We extracted the cirsimaritin from leaves of C. setidum Nakai, and stem and flower of C. pendulum Fisch (Table 1). In fresh tissues, flowers of C. pendulum Fisch had highest amount of cirsimaritin, but the cirsimaritin content in the leaf of C. setidens Nakai and C. pendulum Fisch were not different as amount as 0.019 ± 0.000 mg/g in dried tissues. Stem tissues of C. pendulum showed least amount of cirsimaritin in both fresh and dried tissues.
Table 1.
Cirsimaritin content in leaves, stems, and flowers of Cirsium species
RNAseq results
We obtained 62.5 million raw reads with 6,311Mbp (million base pair), which were trimmed into 60.4 million reads with 6,046 Mbp (Table 2). After removed the redundant sequences (> 98%) from the trimmed results, the number of reads over 240 nucleotides was 42,250 with 44.43 Mbp. Detailed RNAseq result is shown in Table 3.
Table 2.
The pre-processing results of the transcriptomes of leaf tissues of C. setidens Nakai
| Raw Reads | Trimmed data | Retained Rate | ||
| No. Reads | Total length (nt) | No Reads | Total length (nt)) | |
| 62,491,880 | 6,311,679,880 | 60,377,360 | 6,046,251,791 | 93.0 (%) |
Table 3.
De novo transcriptome assembly and unigene transcripts of C. setidum transcriptomes
| De novo transcriptome assemble | Unigene transcripts |
| No of total contig | 40,250 |
| Total length (nt) | 44,439,694 |
| Maximum length (nt) | 15,615 |
| N50 | 1,561 |
Functional annotation
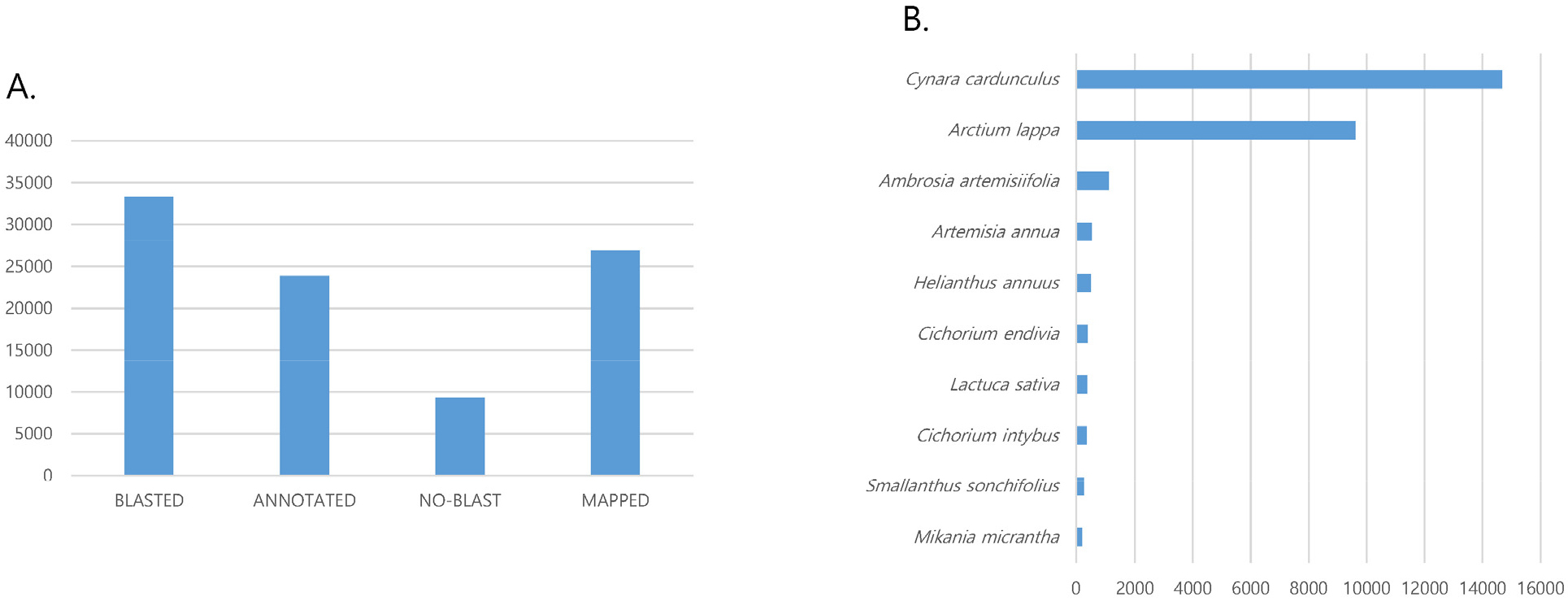
For functional annotation, we queried the sequences of the 42,668 transcripts BLASTX search against non-redundant nucleotide sequences (Nt) and non-redundant protein sequence (Nr) databases in NCBI. Of the obtained transcripts, 71.8% of the transcripts were blasted in the NCBI database and the number of function annotated transcripts was 23,870 (Fig. 1a). The list of the annotated transcripts is available in the http://nbitglobal. com/cirsium. High numbers of C. setidens transcripts were matched with the transcripts of Cyanra cardunculus (14,674) and Articum lappa (9,614). Transcripts of Helianthus annus, Artemisia annua, and Ambrosia artemisiifolia were also matched more than 500 transcripts with those of C. setidens (Fig. 1b).
GO and KEGG analyses
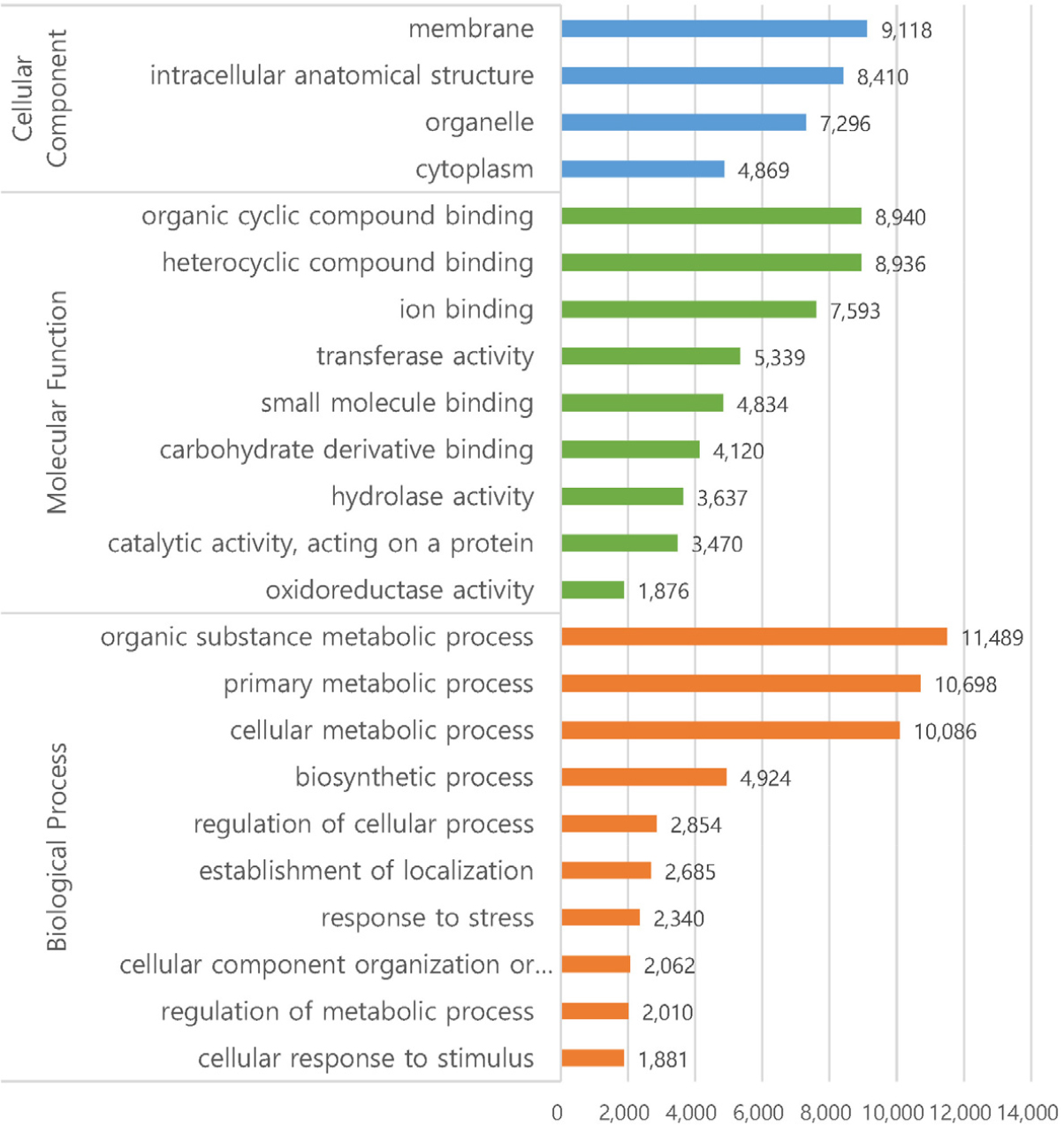
In GO analysis, the 42,668 transcripts were categorized into three functional categories; cellular component, molecular function, and biological process (Fig. 2). In cellular function, four sub-functions were recognized such as membrane, intracellular anatomical structure, organelle, and cytoplasm. In molecular functions, four sub-functions were with over 5000 transcripts such as organic cyclic compound binding, heterocyclic compound binding, ion binding, and transferase activity. In biological process, three sub-functions were predominant with over 10,000 transcripts such as organic substance metabolic process, primary metabolic process, cellular metabolic process, and biosynthetic process.
In KEGG analysis, over 700 transcripts were involved in the purine metabolism (772 transcripts) and thiamine metabolism (725 transcripts), respectively. There were 157, 72 and 62 transcripts in the phenylpropanoid biosynthesis, tyrosine metabolism and phenylalanine metabolism, respectively. The KEGG annotation result is available at http://nbitglobal.com/cirsium.
Isolation of genes in the cirsimaritin biosynthesis
Fig. 3 shows the cirsimaritin biosynthesis pathway which starts from either phenylalanine or tyrosine via stepwise enzyme mediated reactions to lead synthesis of cirsimaritin (Berim and Gang, 2016). We found genes for all enzymes in the pathway from the transcriptome of C. setidens (Table 4). In the annotation, the same gene was annotated to either phenylalanine ammonia (PAL) or tyrosine ammonia lyase (TAL). The number of copies varied from one in trans-cinnamic acid 4-hydroxylase (C4H), flavone 7-O-methyltransferase (F7OMT), and flavone-6-hydroxylase (F6H) to six copies in chalcone synthase (CHS). The genes for cirsimaritin biosynthesis were reported previously by Park et al. (2020) and found out that some genes in our analysis were identical to the genes in their report, but our result revealed extra copies in most genes. Sequences of the enzymes in the pathway are listed at the end of the manuscript.
Table 4.
Cirsimaritin biosynthesis genes in Cirsium setidens Nakai
Note: The number in bracket is the number of copies that are identical to the genes reported by Park et al. (2020).
Phylogenetic analysis
Chalcone synthase (CHS) and chalcone isomerase (CHI) are key enzymes in the flavonoid synthesis. Various kinds of flavonoid-O-methyltransferases play key roles in the flavonoid synthesis (Park et al., 2020) and two flavonoid-O-methyltransferasess, flavonoid-7-O-methyltransferase (F7OMT) and flavonoid-6-O-mehtytransferase (cisimaritin synthase, CRS) are involved in the cirsimaritin biosynthesis. Thus, phylogenetic analyses were carried out with the CHS and CHI.
We downloaded all copies of CHS and CHI from Cynara cardunculus (artichoke thistle), Helianthus annus (sunflower), Silybum marianum (milk thistle) and Glycine max (soybean) in the NCBI protein database. We selected these species because their transcriptomes were highly matched with the transcriptomes of C. setidens. The G. max was selected because soybean was known to contain high amount isoflavones (Wang et al., 2013) and the genes for isoflavone synthesis were well characterized (Gutierrez-Gonzalez et al., 2009). We selected only representative copies by eliminating the duplicated identical proteins in each species using “Identical protein groups” option in the NCBI and ClustalW2 multiple sequence alignment program (https://www.ebi.ac.uk/Tools/ msa/clustalo/).
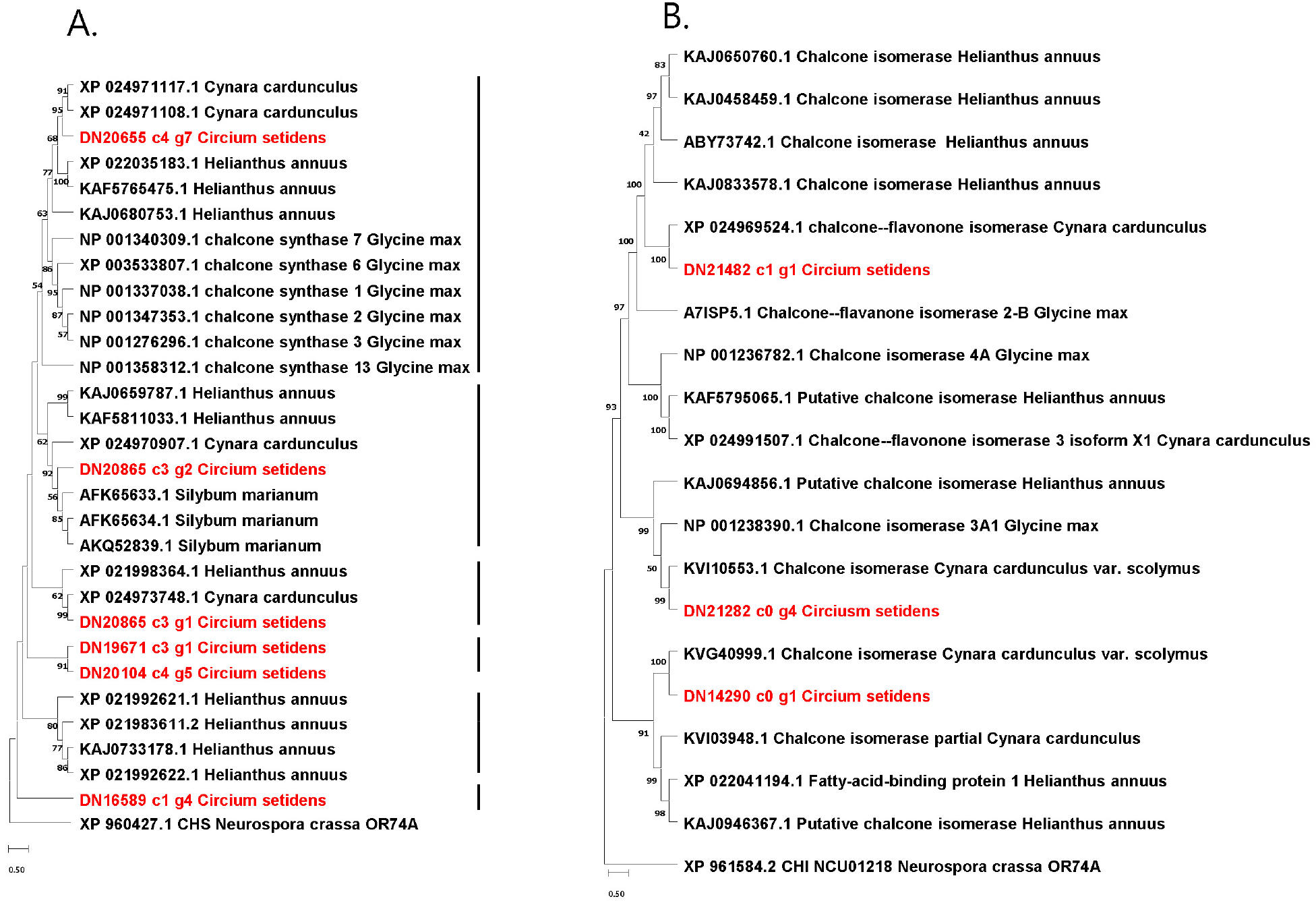
We analyzed 33 CHS proteins that consist of 6, 4, 12, 6, 4, and 1 from C. setidens, C. cardunculus, H. annum, G. max, S. marinum, Neurospora crassa, respectively (Fig. 4A). The CHS of N. crassa, a filamentous fungus, was placed in an out-group in the phylogenetic analysis. The rest 32 CHS proteins were divided into six sub-clades in which the subclade VI formed an out-group clade with a sole CHS of DN16589 c1g4 of C. setidens. The subclade IV contains only two CHS protein of C. setidens with high bootstrap value. The three remaining CHS proteins from C. setidens were fell into one of each subclade I (DN20655 c4 g7), II (DN20865 c3 g2), and III (DN20865 c3 g2). We identified six copies of CHS in C. setidens in which three (DN20655 c4g7, DN20865c3g1, DN16589 c1g4) revealed to be the ones reported by Park et al. (2020), but the other three (DN20865 c3 g2, DN19671 c3 g1, DN20104 c4g5) were not present in their analysis.
We analyzed 20 CHI proteins such as 3, 5, 8, 3 and 1 from C. setidens, C. cardunculus, H. annus, G. max and N. crassa (Fig. 4B). As expected the CHS of N. crassa formed an out-group in the phylogenetic tree. The rest 19 CHI were divided into four clades in which the three CHI of C. setidens were placed into clade I (DN21482 c1 gi), III (DN21282 c0 g4), and IV DN14290 c0 g1. Of the three CHI proteins in our analysis, one (DN21482 c1 g1) was present in the report of Park et al. (2020), but the other two (DN21282 c0g4, DN14290 c0 g1) were noble report in C. setidens.
Discussion
Circium is a large genus that distribute worldwide with about 250 species and ten species have been identified in Korea, including C. setidens Nakai and C. pendulum Fisch that are research subjects of the the current study (Lee, 2002; Song and Kim, 2007). Extracts of the C. setidens have been used for folk medicine to treat various illness such as diabetes, inflammatory symptoms, breast cancer, and other various illnesses (Lai et al., 2014; Park et al., 2017; Wagle et al., 2019). Cirsimaritin is a major bioactive compound in the genus Cirsium. Although several reports are available on the cirsimaritin in C. japonicum (Benali et al., 2022; Lee et al., 2017; Park et al., 2020), it has not been characterized in the C. setidens Nakai and C. pendulum Fisch. We quantified the contents of cirsimaritin from leaves of C. setidum Nakai, and stem and flower of C. pendulum Fisch. Park et al., (2020) reported cirsimaritin was present highest in the leaf as much as 16.55 mg/g, dry weight in C. japomicum. However, we did not obtain this much cirsimaritin in our analysis in C. setidens Nakai and C. pendulum Fisch, implying that cirsimaritin may not present as much as high in C. japonium in the latter two species.
In BLASTX search revealed about 71.8% of the transcriptomes of C. setidens matched in the public database in NCBI in our analysis, which means that 28.2% of the transcripts were unknown their functions. Similar results were obtained in Berberis koreana that is a medicinal plants producing an alkaloid berberine (Roy et al., 2021) and in terpenoids producing Euphorbia maculata (Jeon et al., 2022). Except of the top two species of C. cardunculus and A. lappa, number of matching transcripts were less than 1000 in BLASTX analysis. The C. cardunculus and A. lappa belong the same subtribe Arctiinae in the Asteraceae family (Herrando-Moraira et al., 2019). Thus, the transcriptome sequences may be differentiated more in the subtribe Arctiinae to limit sequencing matching in the BLASTX.
High number of organic cyclic compound binding and intercyclic compound binding in the molecular function and metabolic processes in the biological process in GO analysis is interpreted that the C. japonicum contains many secondary metabolites (Benali et al., 2022; Lee et al., 2017). The cyclic and intercyclic binding proteins as well as the proteins in the metabolic processes may function in the biosynthetic synthesis of the secondary metabolites in C. japonicum.
The genes in the cirsimaritin biosynthesis in our analysis were also reported in a previous study in C. japonicum (Park et al., 2020). However, we identified extra copies in each gene in PAL, 4CL, CHS, CHI, and FNS so that these extra copies are noble in our report in C. setidens. The genes of flavone 7-O-methyltransferase (F7OMT), flavone-6-hydroxylase (F6H) and cirsimaritin synthase (CRS) in our analysis were not reported previously in the genus Cirsium. Gene duplication and functional differentiation are common among the genes encoding secondary metabolites (Ober, 2005; Roy et al., 2021). We also observed multiple copies of the genes encoding the flavonoid synthesis pathway in the current study.
CHS and CHI are key enzymes for various flavonoids in plants so that we further analyzed these two enzymes. We identified 6 and 3 copies of CHS and CHI, respectively, from the transcriptomes of C. setidens. Flavonoids have played indispensable roles in embryophytes (land plants) after colonizing into the lands by providing UV protection, plant defense, and regulatory roles of many genes. More than 6,900 flavonoids with different structures were identified in land plants (Mouradov and Spangenberg, 2014). Enzymes involved in plant metabolism have catalytic promiscuities to catalyze reactions coincidently other than those resctions for evolved (Waki et al., 2020). CHS belongs a broad class family of polyketide synthase enzymes (PKS), known as type III PKS (Abe and Morita, 2010), and CHS is an example of catalytic promiscuity and CHI rectifies the promiscuous CHS activity to ensure flavonoid production by mediating the bidirectional reaction between chalcone and flavone (Jez et al., 2000). CHS is a ubiquitous enzyme in higher plants and mediates the catalyzing reaction of the first step for biosynthesis of various flavonoids that are important plant secondary metabolites (Tohge et al., 2007). CHS converts the malonyl-CoA and 4-coumaroyl-CoA to narignenin chalcone, CoA, and CO2. The six copies of CHS in C. setidens were placed in different clades in clade I, II, III, and V with CHS proteins from other species in the phylogenetic tree, alluding that the CHS genes were differentiated independently in C. setidens. The two copies in the clade IV (DN19671 c3 g1 and DN20104 c4 g5) might have duplicated within the genome of C. setidens.
Chalcone isomerase (CHI) is also called as chalcone-flavone isomerase because it participates in flavonoid biosynthesis (Moustafa and Wong, 1967). Morita et al.(2014) isolated a chalcone isomerase like (CHIL) enzyme from morning glory. Although CHIL lacks the CHI activity, CHIL-mediated flavonoid production was confirmed in various plants in their study. Moreover, it was claimed that the CHIL had evolved from fatty-acid binding protein (Nagaki, 2012). It was proposed that CHIL interacts with CHS and CHI to serve as an activator of them in Arabidopsis thaliana (Jiang et al., 2015). Of the three CHI identified in our study, DN21482 c1g1 was CHI, but DN14290 c0g1 and DN21282 c0g4 were identified as CHIL in blast analysis. However, but the DN14290 c0g1 and DN21282 c0g4 showed very close relationship with chalcone-flavone isomerase or chalcone isomerase of Cynara cardunculus (Fig. 4). Thus, sequence similarity analysis was conducted with the CHIL of C. setidens and CHI C. cardunculus (Fig. 5) and the results showed very high similarity in sequences except of the variable regions in N-terminal or C-terminal regions. Thus, annotation of CHI and CHIL are unequivocal, which might be limit in large scale annotation in genomics without biochemical verification.
One notable feature is the high sequence similarity between the proteins of cirsimaritin biosynthesis between C. setidens and C. cardunculus. In functional annotation, the transcriptomes of C. setidens had highest number of sequence matching with the transcriptome of C. cardunculus. In our phylogenetic analysis of CHS and CHI, seven of the eight proteins of C. setidens showed deepest branch clade with the proteins of C. cardunculus. The common name of the C. cardunculus is cardoon and also called as artichoke thistle. It has been used as folk medicine in Mediterranean region since Roman period (Sonnante et al., 2007). It was known that the cardoon contains variety of bioactive compounds including flavonoids such as naringenin and apigenin, (Silva et al., 2022). Both genera Circium and Cynara belong to the family Asteraceae which is one of the large plant family to contain about 10% of all extant angiosperm species with 250,000 ~ 300,000 species (Mandel et al., 2019). The genera Circium and Cynara are in the same subfamily Carduodeae, tribe Cardueae, and subtribe Carduinae (Herrando-Moraira et al., 2019), thus it is not surprising the close phylogenetic relationships between genes from both species.