Introduction
Materials and Methods
Sample collection and virus identification
Host-response experiment
Sequence analysis and phylogenetic analysis
Results and Discussion
Summary
Introduction
Atractylodes macrocephala is a perennial herbaceous plant belonging to the genus Atractylodes within the Asteraceae family, primarily growing in mountainous regions or dry grasslands. The rhizomes of Atractylodes plants and A. macrocephala have long been used as medicinal ingredients in traditional Korean medicine, particularly for treating various digestive disorders such as indigestion and gastritis (Zhu et al., 2018). Atractylodes macrocephala is a species introduced from China, known for its rapid growth, high yield, and excellent seed production, making it advantageous for cultivation and seed propagation. However, it has been reported to be more susceptible to disease compared to other plants in the genus Atractylodes. To date, the major viruses reported in A. macrocephala include Atractylodes mild mottle virus (AMMV), Atractylodes mosaic virus (AtrMoV), Broad bean wilt virus 2 (BBWV2), Pecan mosaic-associated virus (PMaV), and Cucumber mosaic virus (CMV) (Kwon et al., 2019; Lim et al., 2015; Niu et al., 2015, 2018; Zhao et al., 2015).
CMV is an economically significant plant virus with a broad host range infecting over 1,200 plant species worldwide, causing various symptoms, including mosaic and yellowing. The CMV genome consists of three single-stranded RNA segments, with RNA3 encoding the movement protein (MP) and coat protein (CP). These two proteins are known to play a crucial role in disease manifestation and host range determination (Mochizuki and Ohki, 2012). Therefore, comparative analysis of CMV strains isolated from specific host plants holds significant importance for understanding the virus's pathogenicity, host adaptability, and genetic diversity.
Although three CMV isolates separated from A. macrocephala in China and South Korea have been previously reported, systematic comparative studies analyzing their biological responses and molecular diversities are lacking (Kwon et al., 2019; Song et al., 2020). In this study, we analyzed the biological and molecular characteristics of CMV-Atm, newly isolated from A. macrocephala in the Chuncheon area of Gangwon Province, by comparing it with three previously isolated strains. Furthermore, we attempted to elucidate the genetic diversity of CMVs within the host plant, A. macrocephala, and the molecular characteristics associated with host adaptation by comparing RNA3 sequences.
Materials and Methods
Sample collection and virus identification
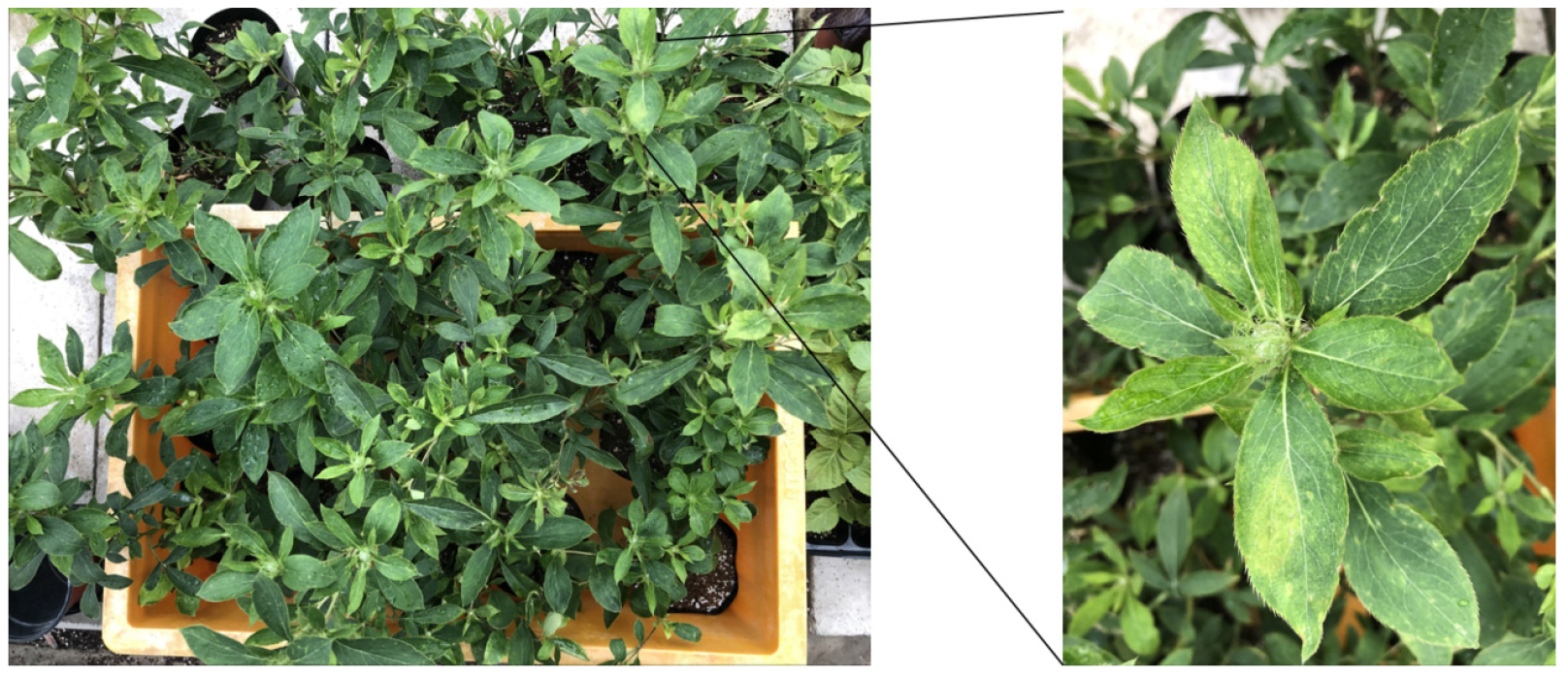
In 2020, A. macrocephala exhibiting distortion symptoms was collected in the Chuncheon area of Gangwon Province (Fig. 1). Total RNA was extracted from symptomatic leaves using the BCSTM Plant RNA Prep Kit (Biocube System Inc., Suwon, South Korea) to determine whether viral infection was present. The extracted total RNA was reverse transcribed into cDNA using M-MLV reverse transcriptase (Promega, Madison, WI, USA), with the 3’-terminal reverse primer used for detection. Reverse transcription-polymerase chain reaction (RT-PCR) was performed using rTaq polymerase (Takara, Japan) with the synthesized cDNA as a template.
The reverse transcription reaction conditions were set at 42°C for 1 h and 92°C for 5 min. PCR amplification was performed under the following conditions: 35 cycles of 30 s at 95°C, 30 s at 52°C, and 1 min at 72°C. Virus detection was performed using genus-specific primer sets for seven virus genera (Ilarvirus, Potyvirus, Fabavirus, Tospovirus, Potexvirus, Tobamovirus, and Cucumovirus), along with AMMV, AtrMoV, BBWV2, and PMaV-specific primers reported for A. macrocephala (Table 1). RT-PCR products were electrophoresed on a 1% agarose gel for 30 min, and bands were visualized under UV light.
Table 1
Primer list used for virus detection in A. macrocephala
| Virus | Primer name | Sequence (5’-3’)1) | Size (bp) | Reference |
| AMMV | AMMV-CP-F | TGGCTGACATACTTGCAGAAGC | 625 | This study |
| AMMV-CP-R | AGTGCCCTTGCTCACTGCA | |||
| AtrMoV | AtrMV-CP-F | ATGCCGCCCAAGCCTGAT | 947 | This study |
| AtrMV-CP-R | TTACTGAAGACTCCGGTTATTCGC | |||
| BBWV2 | BBWV2-506-F | GGTGAGCAGTTTGTCAGAAGT | 506 | Choi et al. (2019) |
| BBWV2-506-R | CCAGATAATGCATATTCCACC | |||
| PMaV | PMaV-CP-F | CCACCGCTAACTAGCGCT | 750 | This study |
| PMaV-CP-R | AACATCATCGGTGGTGTGTC | |||
| Ilarvirus | Ilar-5’ | GCNGGWTGYGGDAARWCNAC | 309 | Untiveros et al. (2010) |
| Ilar-3’ | AMDGGWAYYTGYTYNGTRTCACC | |||
| Fabavirus | Faba-genus-5' | AAATATTAAAACAAACAGCTTTCGTT | 390 | Ferrer et al. (2007) |
| Faba-genus-3' | TTCAAAGCTCGTGCCATITYATTKGC | |||
| Tospovirus | Tospovirus-5' | TCIRDICKIYKRAAICTCMSRTC | 450 | Okuda and Hanada (2001) |
| Tospovirus-3’ | GGGGGAGAGCAATYGWGTCA | |||
| Potexvirus | Potexvirus-5 | CAYCARCARGCMAARGAYGA | 580 | Van der Vlugt and Berendsen (2002) |
| Potexvirus-3 | AGCATRGCISCRTCYTG | |||
| Potyvirus | Poty-CI-For | GGIVVIGTIGGIWSIGGIAARTCIAC | 700 | Ha et al. (2008) |
| Poty-CI-Rev | ACICCRTTYTCDATDATRTTIGTIGC | |||
| Tobamovirus | TobamodF | TKGAYGGNGTBCCNGGNTGYGG | 880 | Li et al. (2018) |
| TobamodR | ACNGAVTBNABCTGTAATTGCTAT | |||
| Cucumovirus | CPT-All-F | YASYTTTDRGGTTCAATTCC | 950 | Choi et al. (1999) |
| CPT-All-R | GACTGACCATTTTAGCCG |
Host-response experiment
Mechanical inoculation was performed on 10 host plant species, including Nicotiana benthamiana (Table 2), to examine the host response of CMV-Atm. Leaves from A. macrocephala plant showing symptoms (1 g) were crushed in a mortar with 1 mL of 0.01 M phosphate buffer (pH 7.2) and used as an inoculum. The prepared extract was rubbed onto the leaf surface of the host plant treated with carborundum for inoculation. After the inoculation, plants were cultivated in a growth chamber maintained at 25°C–27°C and 60% relative humidity, with a 16-h light/8-h dark photoperiod.
Table 2
Host response to CMV-Atm
| Family | Host plants | Symptoms of the leaves1) | ||
| JC2) | Atm | Fny3) | ||
| Solanaceae | Nicotiana benthamiana | -/sM | -/M | -/M |
| N. tabacum cv. Xanthi nc | nt | NS/M | -/M | |
| N. glutinosa | -/sM | NS/M | -/M | |
| Capsicum annuum cv. Cheongyang | nt | -/M | -/M | |
| C. annuum cv. Baerotta | nt | -/- | -/- | |
| C. annuum cv. Bukang | nt | -/- | -/- | |
| Chenopodiaceae | Chenopodium quinoa | nt | NS/- | NS/- |
| C. amaranticolor | nt | NS/- | NS/- | |
| Cucurbitaceae | Cucumis sativus cv. Daesun | nt | -/CS | -/M |
| Cucurbita pepo cv. Choigobong | nt | NS/CS | -/M | |
1)Inoculated/upper leaves; sM, severe mosaic; M, mosaic; NS, necrotic spot; CS, chlorotic spot; -, no infection; nt, not tested.
CMV-JC2) and CMV-Fny3) were used as a control (Banik et al., 1983; Kwon et al., 2019)
Sequence analysis and phylogenetic analysis
RT-PCR was performed using the CMV-I-RNA3-F primer (5’-GTA ATC TTA CCA CTG TGT GTG-3’) and the SSVK3-3’-BamHI primer (5’-GCG GAT CCT GGT CTC CTT TGG AAG CCC-3’) to amplify the entire sequence of CMV-Atm RNA3 (Lee, 2007). The amplified PCR products were cloned into the pGEM T-easy TA cloning vector system (Promega, USA) and subsequently sequenced by Macrogen (Seoul, South Korea). The obtained nucleotide sequences were analyzed for homology using the BLASTn program from NCBI and MEGA10 software.
The phylogenetic analysis of CMV-Atm included sequences from 16 CMV isolates, with peanut stunt virus (PSV-ER) as the outgroup. The nucleotide sequences of other CMV isolates were obtained from the GenBank database. The sequences were subjected to multiple alignment using ClustalW in MEGA10 software. A phylogenetic tree was constructed using the maximum likelihood method with 1,000 bootstrap iterations (Fig. 2). Furthermore, the amino acid sequences of the coat protein from CMV-Atm and the existing A. macrocephala-derived CMV isolates were aligned and compared using the BioEdit program (Fig. 3).
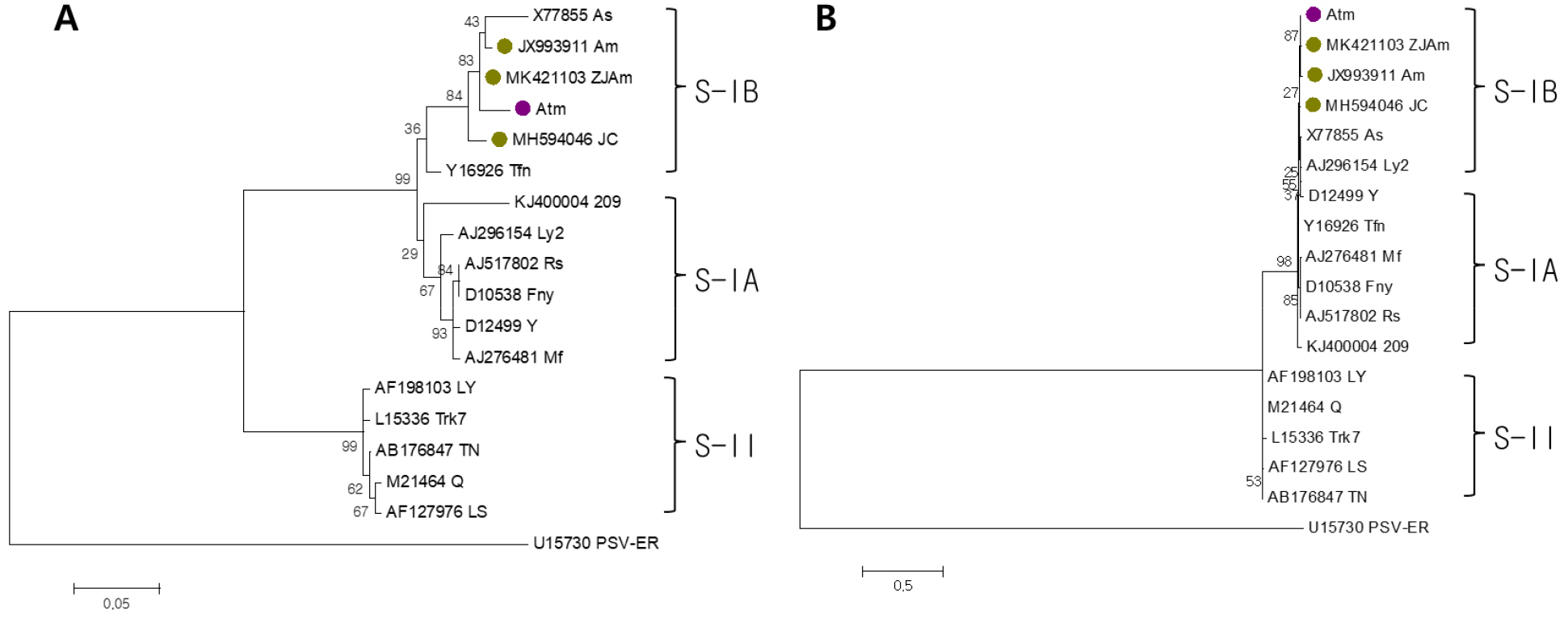
Fig. 2.
Phylogenetic analysis based on the amino acid sequences of movement protein (A) and coat protein (B) of CMV-Atm, together with other CMV isolates including those derived from A. macrocephala. Phylogenetic analysis was conducted using the maximum likelihood method with 1,000 bootstrap replications implemented in MEGA 10 after multiple alignment by ClustalW. CMV isolates were classified into three subgroups, and peanut stunt virus (PSV-ER) was used as outgroup. The sequence of other CMV isolates were obtained from the GenBank database. The purple circle indicates CMV-Atm identified in this study, and the green circles represent previously reported CMV isolates from A. macrocephala.
Results and Discussion
We tested whether virus infection was present in A. macrocephala exhibiting distortion symptoms in the Chuncheon region. After extracting total RNA from leaves exhibiting symptoms, RT-PCR testing was performed for previously reported viruses, i.e., AMMV, AtrMoV, BBWV2, and PMaV (Table 1). Supplementary tests using genus-specific primer sets for seven viral genera, i.e., Ilarvirus, Potyvirus, Fabavirus, Tospovirus, Potexvirus, Tobamovirus, and Cucumovirus., were also conducted to determine the potential infection of other viruses. RT-PCR results showed that only the test primer set within the Cucumovirus genus produced an amplification product of the expected size (approximately 950 bp) (data not shown). Sequence analysis of the amplified product confirmed that A. macrocephala was infected with CMV, and this isolate was designated CMV-Atm. The acquired nucleotide sequence showed 99.85% of sequence homology with that of the CMV-Ad71 isolated in Angelica dahurica (GenBank accession no. PV939598.1).
To examine the biological characteristics of CMV-Atm, mechanical inoculation was performed on 10 host plant species, including Nicotiana benthamiana (Table 2). Three days after inoculation, necrotic spots appeared on the inoculated leaves of N. tabacum cv. Xanthi nc, and N. glutinosa. Ten days after inoculation, typical mosaic symptoms were observed on the upper leaves of all tobacco plants. In Capsicum annuum, mosaic symptoms were found only in the susceptible cultivar 'Cheongyang'. In Chenopodium quinoa and C. amaranticolor, necrotic spots were observed only on the inoculated leaves. Additionally, yellow spots appeared on the upper leaves of the Cucurbit crops, including cucumber and zucchini.
When comparing these host responses to those of CMV-JC, previously isolated from A. macrocephala, and CMV-Fny, a representative CMV isolate, CMV-JC and CMV-Fny induced mosaic symptoms on the upper leaves of N. benthamiana and N. glutinosa. In contrast, CMV-Atm caused necrotic spots on the inoculated leaves (Kwon et al., 2019). Unlike CMV-Fny, which induced upper leaf mosaic symptoms in cucurbit crops, CMV-Atm caused yellow spots. These results indicated that CMV-Atm possesses distinct pathogenicity patterns in certain hosts (tobacco plants), while having a host range similar to that of typical CMV strains. Furthermore, CMV-Atm failed to infect or induce disease symptoms in resistant pepper varieties, indicating its inability to overcome resistance in these varieties.
To analyze the genetic characteristics of CMV-Atm, we performed phylogenetic analysis on the movement protein (MP) and coat protein (CP) genes encoded by RNA3. CMV-Atm was found to belong to the IB group within subgroup I in terms of both MP and CP (Fig. 2). This was similar to previously reported A. macrocephala-derived CMV strains. Meanwhile, CMV-Atm showed a higher association with the Chinese isolates CMV-ZJAm and CMV-Am than with the Korean isolate CMV-JC.
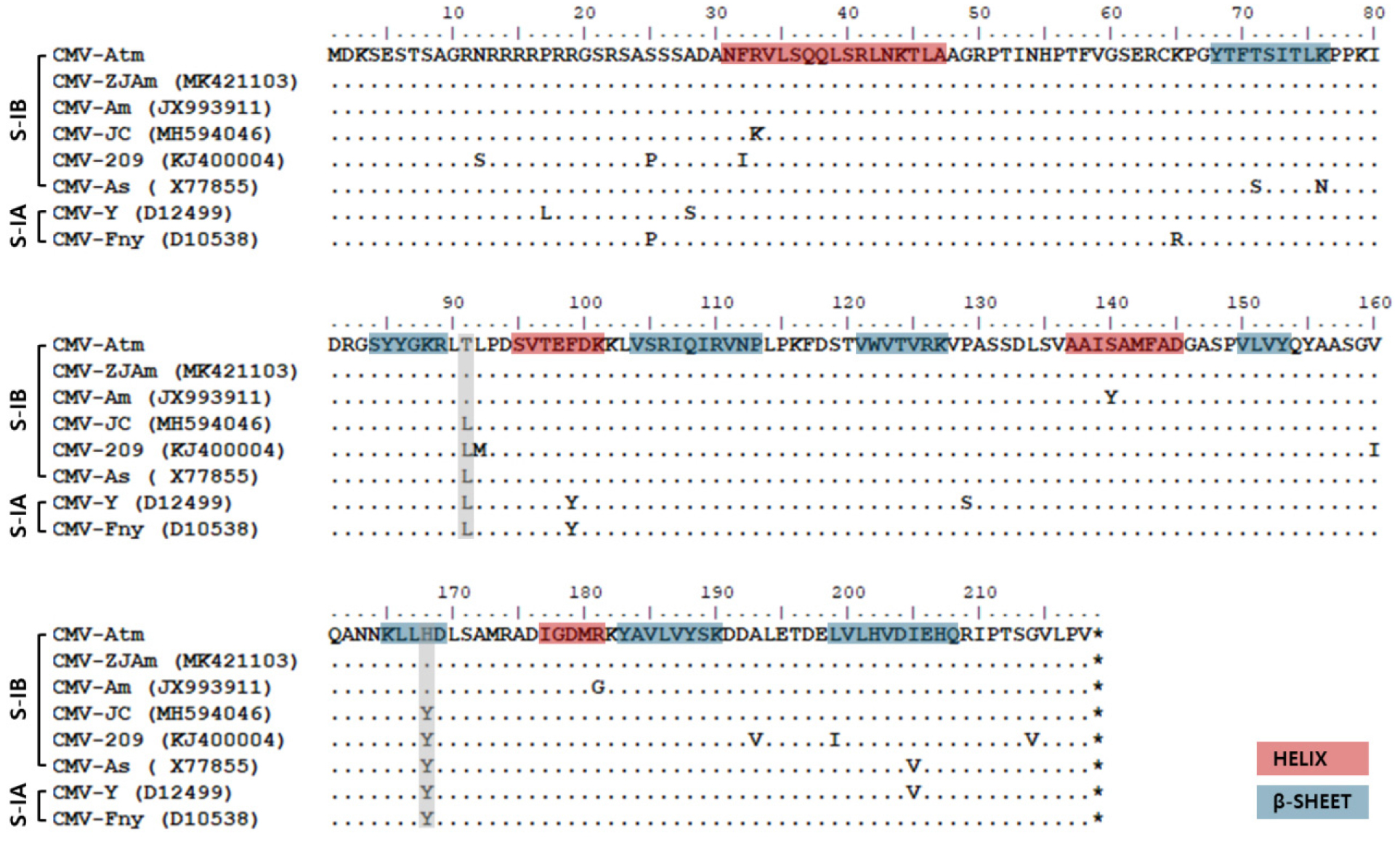
Additionally, we compared the CP amino acid sequences of CMV-Atm with those of three previously separated CMV isolates from A. macrocephala and various major isolates within subgroup I (Fig. 3). CMV-Atm exhibited amino acid sequences nearly identical to those of existing A. macrocephala-derived isolates. Particularly, amino acids at positions 91 and 168 of CMV-Atm were specific to those isolates from A. macrocephala, distinguishing them from other isolates belonging to subgroups IA and IB. The helix and β-sheet regions of the coat protein are known to be critical for the protein's structural stability and folding (Smith et al., 2000). In the CMV-Atm sequence, slight variation was observed in this region, suggesting that the CP of A. macrocephala–derived CMV isolates is highly conserved.
This study confirmed that CMV-Atm exhibits high phylogenetic similarity to existing isolates derived from A. macrocephala, yet displays distinct disease symptoms in certain host responses. This suggests that within A. macrocephala host, CMV maintains overall conservative evolution while accumulating host adaptation and variation at the level of pathogenicity-related genes or proteins.
Summary
Atractylodes macrocephala is a medicinal crop belonging to the Atractylodes genus, widely cultivated in China and South Korea. To date, five viruses have been reported to infect A. macrocephala. In this study, samples of A. macrocephala exhibiting distortion symptoms were collected in Chuncheon, Gangwon Province. RT-PCR testing was performed targeting five viruses reported on this host plant and seven virus genera commonly occurring in South Korea. As a result, an amplification product of the expected size was detected using a Cucumovirus genus-specific primer set. Sequence analysis confirmed infection with cucumber mosaic virus (CMV), and this isolate was designated CMV-Atm. Previously, CMV isolates identified in A. macrocephala have been reported in China and South Korea. Therefore, this study compared the biological and molecular characteristics of the newly isolated CMV-Atm with existing A. macrocephala–derived CMV isolates. CMV-Atm exhibited differences in pathogenicity and host response compared with existing isolates, yet showed high sequence homology with them in phylogenetic analysis. These results suggest that CMV-Atm possesses genetically conserved characteristics with existing CMV isolates derived from A. macrocephala, while exhibiting unique traits in the manifestation of disease symptoms, indicating the possibility of subtle adaptive processes occurring during host-pathogen interactions.